| Show | PDB | Class | SubFamily | Type | SubType | Species | Orthosteric Ligand | Other Ligand(s) | Protein Partners | Resolution | Date | DOI |
| 8ZW0 | A | Lipid | Prostaglandin | DP1 | Homo Sapiens | PGD2 | - | Gs/Beta1/Gamma2 | 2.72 | 2025-05-28 | doi.org/10.1073/pnas.2501902122 |
|
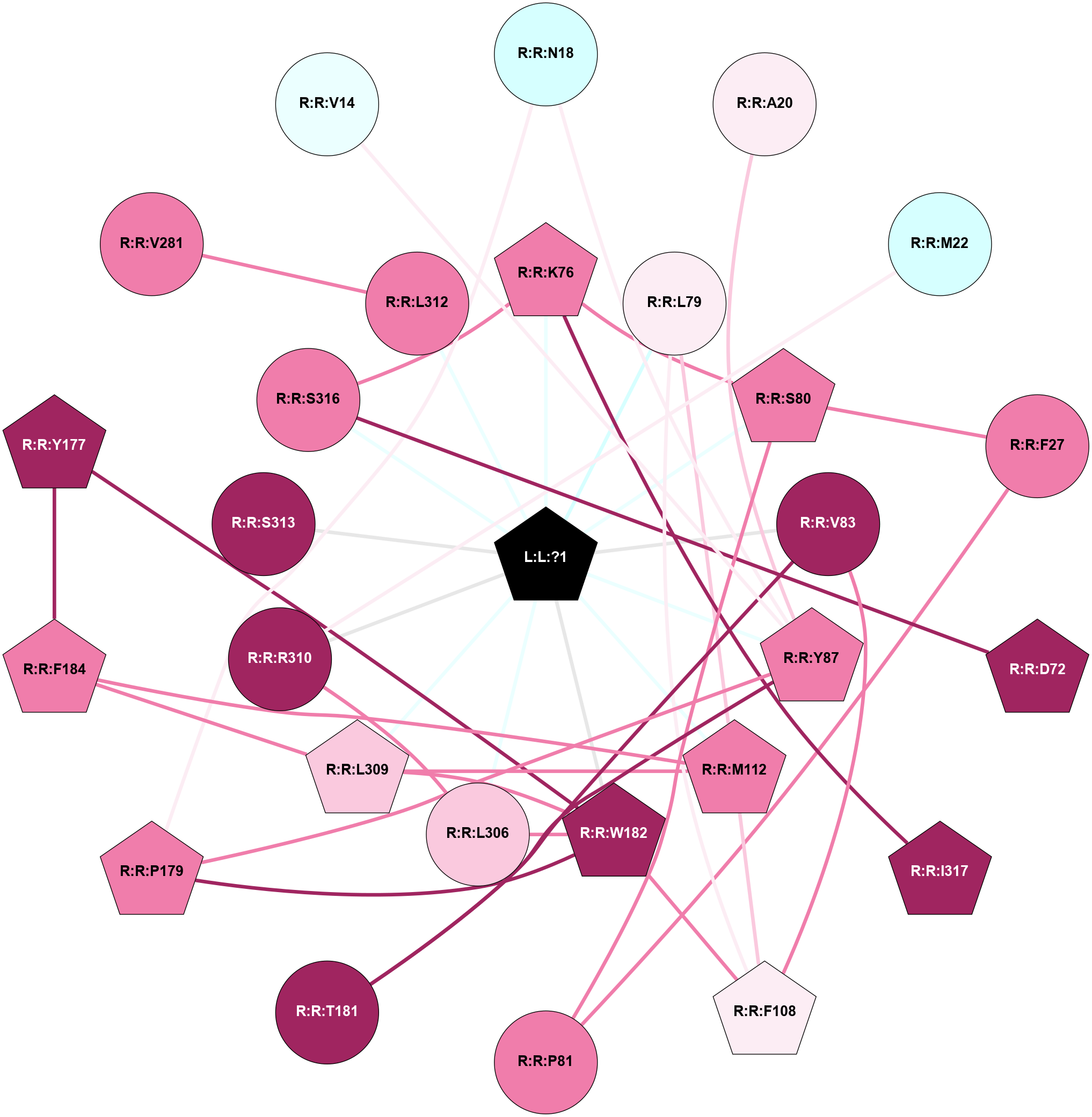
A 2D representation of the interactions of PG2 in 8ZW0
colored according to ConSurf Conservation Grade (See documentation):
(Click to enlarge 🔍) | | Node1 | Node2 | LinkStrength | Comm | IsNode1Hub? | IsNode2Hub? | Node1Cons | Node2Cons | Node1Shell | Node2Shell | | R:R:N18 | R:R:Y87 | 9.3 | 1 | No | Yes | 3 | 8 | 2 | 1 | | R:R:N18 | R:R:P179 | 3.26 | 1 | No | Yes | 3 | 8 | 2 | 2 | | R:R:A20 | R:R:Y87 | 9.34 | 0 | No | Yes | 6 | 8 | 2 | 1 | | R:R:M22 | R:R:R310 | 14.89 | 0 | No | No | 3 | 9 | 2 | 1 | | R:R:F27 | R:R:S80 | 11.89 | 1 | No | Yes | 8 | 8 | 2 | 1 | | R:R:F27 | R:R:P81 | 5.78 | 1 | No | No | 8 | 8 | 2 | 2 | | R:R:D72 | R:R:S316 | 8.83 | 0 | Yes | No | 9 | 8 | 2 | 1 | | R:R:K76 | R:R:S80 | 3.06 | 1 | Yes | Yes | 8 | 8 | 1 | 1 | | R:R:K76 | R:R:S316 | 6.12 | 1 | Yes | No | 8 | 8 | 1 | 1 | | R:R:I317 | R:R:K76 | 4.36 | 0 | Yes | Yes | 9 | 8 | 2 | 1 | | L:L:?1 | R:R:K76 | 13.91 | 1 | Yes | Yes | 0 | 8 | 0 | 1 | | R:R:F108 | R:R:L79 | 12.18 | 1 | Yes | No | 6 | 6 | 2 | 1 | | R:R:L79 | R:R:M112 | 4.24 | 1 | No | Yes | 6 | 8 | 1 | 1 | | L:L:?1 | R:R:L79 | 7.28 | 1 | Yes | No | 0 | 6 | 0 | 1 | | R:R:P81 | R:R:S80 | 3.56 | 1 | No | Yes | 8 | 8 | 2 | 1 | | L:L:?1 | R:R:S80 | 8.89 | 1 | Yes | Yes | 0 | 8 | 0 | 1 | | R:R:F108 | R:R:V83 | 3.93 | 1 | Yes | No | 6 | 9 | 2 | 1 | | R:R:T181 | R:R:V83 | 3.17 | 0 | No | No | 9 | 9 | 2 | 1 | | L:L:?1 | R:R:V83 | 7.84 | 1 | Yes | No | 0 | 9 | 0 | 1 | | R:R:P179 | R:R:Y87 | 4.17 | 1 | Yes | Yes | 8 | 8 | 2 | 1 | | R:R:T181 | R:R:Y87 | 17.48 | 0 | No | Yes | 9 | 8 | 2 | 1 | | L:L:?1 | R:R:Y87 | 6.17 | 1 | Yes | Yes | 0 | 8 | 0 | 1 | | R:R:F108 | R:R:M112 | 3.73 | 1 | Yes | Yes | 6 | 8 | 2 | 1 | | R:R:F108 | R:R:W182 | 3.01 | 1 | Yes | Yes | 6 | 9 | 2 | 1 | | R:R:F184 | R:R:M112 | 8.71 | 1 | Yes | Yes | 8 | 8 | 2 | 1 | | R:R:L309 | R:R:M112 | 2.83 | 1 | Yes | Yes | 7 | 8 | 1 | 1 | | L:L:?1 | R:R:M112 | 9.3 | 1 | Yes | Yes | 0 | 8 | 0 | 1 | | R:R:W182 | R:R:Y177 | 26.04 | 1 | Yes | Yes | 9 | 9 | 1 | 2 | | R:R:F184 | R:R:Y177 | 4.13 | 1 | Yes | Yes | 8 | 9 | 2 | 2 | | R:R:P179 | R:R:W182 | 6.76 | 1 | Yes | Yes | 8 | 9 | 2 | 1 | | R:R:L306 | R:R:W182 | 4.56 | 1 | No | Yes | 7 | 9 | 1 | 1 | | R:R:L309 | R:R:W182 | 3.42 | 1 | Yes | Yes | 7 | 9 | 1 | 1 | | L:L:?1 | R:R:W182 | 14.98 | 1 | Yes | Yes | 0 | 9 | 0 | 1 | | R:R:F184 | R:R:L309 | 7.31 | 1 | Yes | Yes | 8 | 7 | 2 | 1 | | R:R:L312 | R:R:V281 | 5.96 | 0 | No | No | 8 | 8 | 1 | 2 | | R:R:L306 | R:R:R310 | 14.58 | 1 | No | No | 7 | 9 | 1 | 1 | | L:L:?1 | R:R:L306 | 4.55 | 1 | Yes | No | 0 | 7 | 0 | 1 | | L:L:?1 | R:R:L309 | 2.73 | 1 | Yes | Yes | 0 | 7 | 0 | 1 | | L:L:?1 | R:R:R310 | 9.59 | 1 | Yes | No | 0 | 9 | 0 | 1 | | L:L:?1 | R:R:S313 | 9.88 | 1 | Yes | No | 0 | 9 | 0 | 1 | | L:L:?1 | R:R:S316 | 2.96 | 1 | Yes | No | 0 | 8 | 0 | 1 | | R:R:V14 | R:R:Y87 | 2.52 | 0 | No | Yes | 4 | 8 | 2 | 1 | | L:L:?1 | R:R:L312 | 1.82 | 1 | Yes | No | 0 | 8 | 0 | 1 | |
| Statistics | Value |
|---|
| Average Number Of Links | 13.00 | | Average Number Of Links With An Hub | 6.00 | | Average Interaction Strength | 7.68 | | Average Nodes In Shell | 28.00 | | Average Hubs In Shell | 13.00 | | Average Links In Shell | 43.00 | | Average Links Mediated by Hubs In Shell | 38.00 |
| 
Physico-chemical properties of the nodes interacting with this ligand (click to enlarge 🔍) | 
Location of the nodes interacting with this ligand (click to enlarge 🔍) |
|
| Show | PDB | Class | SubFamily | Type | SubType | Species | Orthosteric Ligand | Other Ligand(s) | Protein Partners | Resolution | Date | DOI |
| 9IYB | A | Lipid | Prostanoid | DP2 | Homo Sapiens | PGD2 | A1D5Q | Gi1/Beta1/Gamma2 | 2.82 | 2024-12-04 | doi.org/10.1073/pnas.2403304121 |
|
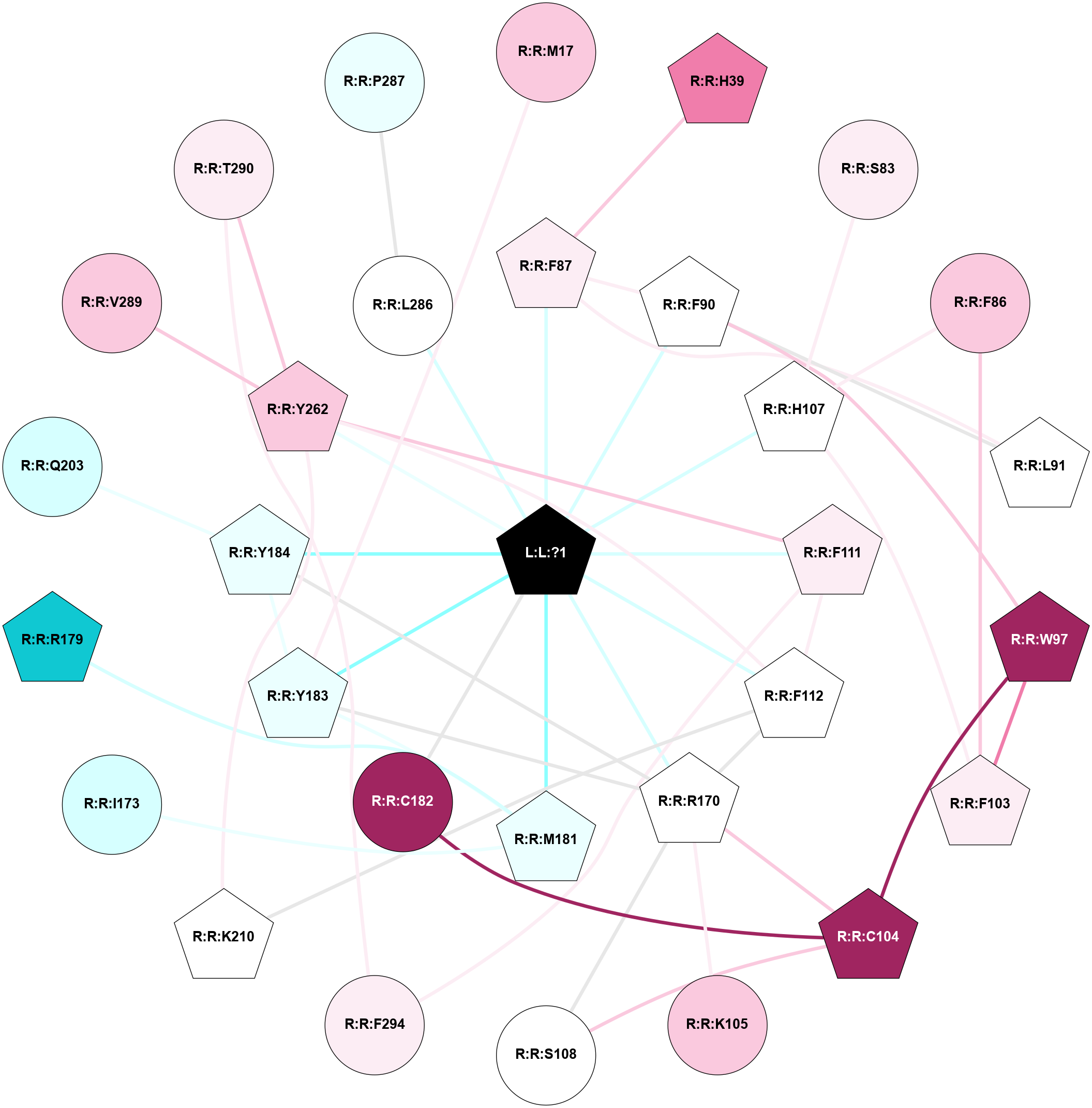
A 2D representation of the interactions of PG2 in 9IYB
colored according to ConSurf Conservation Grade (See documentation):
(Click to enlarge 🔍) | | Node1 | Node2 | LinkStrength | Comm | IsNode1Hub? | IsNode2Hub? | Node1Cons | Node2Cons | Node1Shell | Node2Shell | | L:L:?1 | R:R:F87 | 12.02 | 1 | Yes | Yes | 0 | 6 | 0 | 1 | | L:L:?1 | R:R:F90 | 25.63 | 1 | Yes | Yes | 0 | 5 | 0 | 1 | | L:L:?1 | R:R:H107 | 10.15 | 1 | Yes | Yes | 0 | 5 | 0 | 1 | | L:L:?1 | R:R:F111 | 6.41 | 1 | Yes | Yes | 0 | 6 | 0 | 1 | | L:L:?1 | R:R:F112 | 7.21 | 1 | Yes | Yes | 0 | 5 | 0 | 1 | | L:L:?1 | R:R:R170 | 7.99 | 1 | Yes | Yes | 0 | 5 | 0 | 1 | | L:L:?1 | R:R:M181 | 20.46 | 1 | Yes | Yes | 0 | 4 | 0 | 1 | | L:L:?1 | R:R:C182 | 5.22 | 1 | Yes | No | 0 | 9 | 0 | 1 | | L:L:?1 | R:R:Y183 | 4.63 | 1 | Yes | Yes | 0 | 4 | 0 | 1 | | L:L:?1 | R:R:Y184 | 17.73 | 1 | Yes | Yes | 0 | 4 | 0 | 1 | | L:L:?1 | R:R:Y262 | 3.08 | 1 | Yes | Yes | 0 | 7 | 0 | 1 | | L:L:?1 | R:R:L286 | 3.64 | 1 | Yes | No | 0 | 5 | 0 | 1 | | R:R:M17 | R:R:Y183 | 8.38 | 0 | No | Yes | 7 | 4 | 2 | 1 | | R:R:F87 | R:R:H39 | 3.39 | 1 | Yes | Yes | 6 | 8 | 1 | 2 | | R:R:H107 | R:R:S83 | 9.76 | 1 | Yes | No | 5 | 6 | 1 | 2 | | R:R:F103 | R:R:F86 | 16.08 | 1 | Yes | No | 6 | 7 | 2 | 2 | | R:R:F86 | R:R:H107 | 4.53 | 1 | No | Yes | 7 | 5 | 2 | 1 | | R:R:F87 | R:R:F90 | 3.22 | 1 | Yes | Yes | 6 | 5 | 1 | 1 | | R:R:F87 | R:R:L91 | 12.18 | 1 | Yes | Yes | 6 | 5 | 1 | 2 | | R:R:F90 | R:R:L91 | 3.65 | 1 | Yes | Yes | 5 | 5 | 1 | 2 | | R:R:F90 | R:R:W97 | 9.02 | 1 | Yes | Yes | 5 | 9 | 1 | 2 | | R:R:F103 | R:R:W97 | 6.01 | 1 | Yes | Yes | 6 | 9 | 2 | 2 | | R:R:C104 | R:R:W97 | 3.92 | 1 | Yes | Yes | 9 | 9 | 2 | 2 | | R:R:F103 | R:R:H107 | 4.53 | 1 | Yes | Yes | 6 | 5 | 2 | 1 | | R:R:C104 | R:R:S108 | 3.44 | 1 | Yes | No | 9 | 5 | 2 | 2 | | R:R:C104 | R:R:R170 | 5.57 | 1 | Yes | Yes | 9 | 5 | 2 | 1 | | R:R:C104 | R:R:C182 | 7.28 | 1 | Yes | No | 9 | 9 | 2 | 1 | | R:R:K105 | R:R:R170 | 3.71 | 0 | No | Yes | 7 | 5 | 2 | 1 | | R:R:R170 | R:R:S108 | 15.81 | 1 | Yes | No | 5 | 5 | 1 | 2 | | R:R:F111 | R:R:F112 | 9.65 | 1 | Yes | Yes | 6 | 5 | 1 | 1 | | R:R:F111 | R:R:Y262 | 8.25 | 1 | Yes | Yes | 6 | 7 | 1 | 1 | | R:R:F111 | R:R:F294 | 11.79 | 1 | Yes | No | 6 | 6 | 1 | 2 | | R:R:F112 | R:R:R170 | 5.34 | 1 | Yes | Yes | 5 | 5 | 1 | 1 | | R:R:F112 | R:R:K210 | 4.96 | 1 | Yes | Yes | 5 | 5 | 1 | 2 | | R:R:F112 | R:R:Y262 | 3.09 | 1 | Yes | Yes | 5 | 7 | 1 | 1 | | R:R:R170 | R:R:Y183 | 3.09 | 1 | Yes | Yes | 5 | 4 | 1 | 1 | | R:R:R170 | R:R:Y184 | 3.09 | 1 | Yes | Yes | 5 | 4 | 1 | 1 | | R:R:I173 | R:R:M181 | 4.37 | 1 | No | Yes | 3 | 4 | 2 | 1 | | R:R:M181 | R:R:R179 | 4.96 | 1 | Yes | Yes | 4 | 1 | 1 | 2 | | R:R:M181 | R:R:Y183 | 10.78 | 1 | Yes | Yes | 4 | 4 | 1 | 1 | | R:R:Y183 | R:R:Y184 | 17.87 | 1 | Yes | Yes | 4 | 4 | 1 | 1 | | R:R:Q203 | R:R:Y184 | 15.78 | 0 | No | Yes | 3 | 4 | 2 | 1 | | R:R:K210 | R:R:Y262 | 4.78 | 1 | Yes | Yes | 5 | 7 | 2 | 1 | | R:R:V289 | R:R:Y262 | 3.79 | 0 | No | Yes | 7 | 7 | 2 | 1 | | R:R:T290 | R:R:Y262 | 4.99 | 0 | No | Yes | 6 | 7 | 2 | 1 | | R:R:L286 | R:R:P287 | 3.28 | 0 | No | No | 5 | 4 | 1 | 2 | | R:R:F294 | R:R:T290 | 5.19 | 0 | No | No | 6 | 6 | 2 | 2 | |
| Statistics | Value |
|---|
| Average Number Of Links | 12.00 | | Average Number Of Links With An Hub | 10.00 | | Average Interaction Strength | 10.35 | | Average Nodes In Shell | 31.00 | | Average Hubs In Shell | 18.00 | | Average Links In Shell | 47.00 | | Average Links Mediated by Hubs In Shell | 45.00 |
| 
Physico-chemical properties of the nodes interacting with this ligand (click to enlarge 🔍) | 
Location of the nodes interacting with this ligand (click to enlarge 🔍) |
|
| Show | PDB | Class | SubFamily | Type | SubType | Species | Orthosteric Ligand | Other Ligand(s) | Protein Partners | Resolution | Date | DOI |
| 9E9S | A | Lipid | Prostanoid | DP1 | Homo Sapiens | PGD2 | - | Gs/Beta1/Gamma1 | 2.68 | 2025-09-03 | To be published |
|
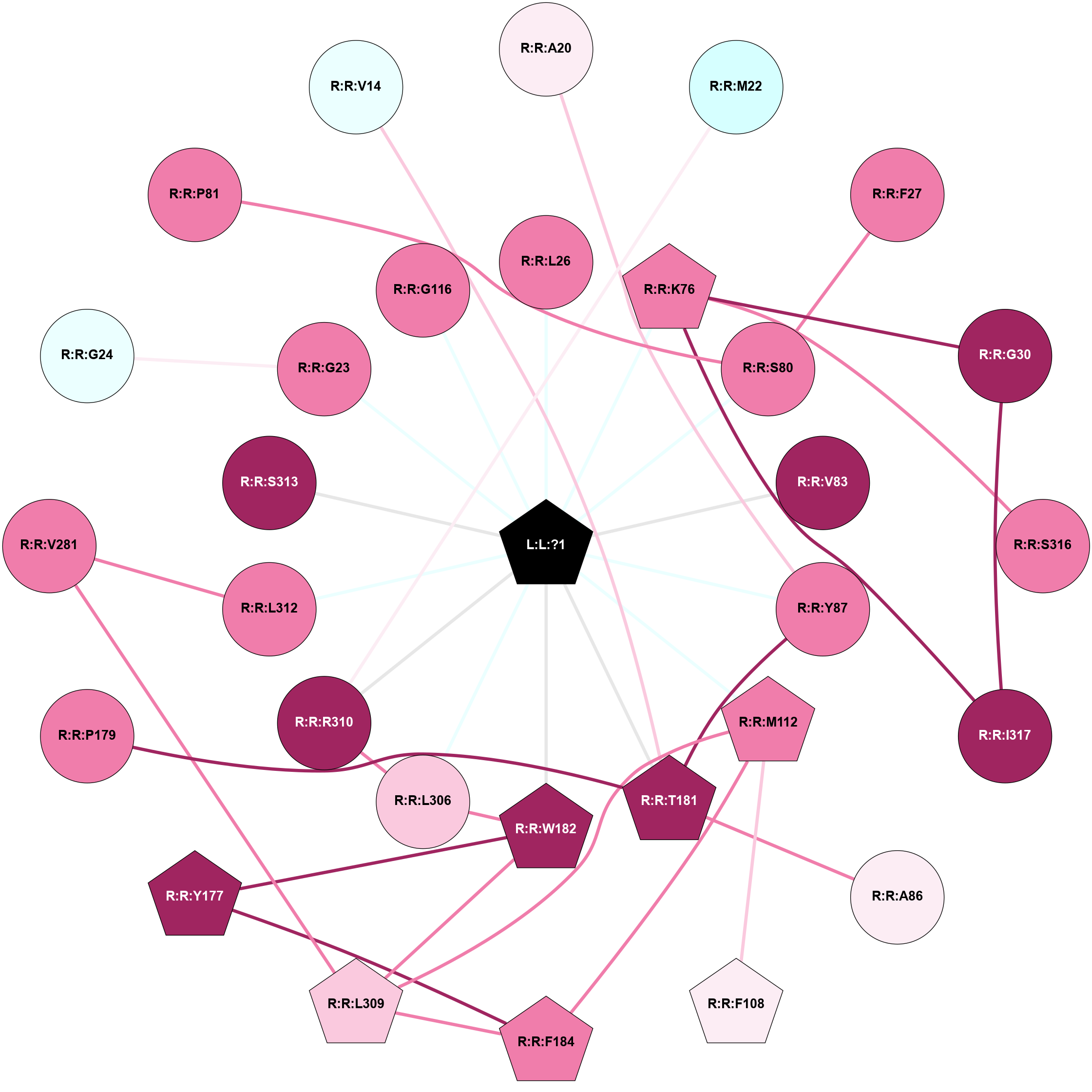
A 2D representation of the interactions of PG2 in 9E9S
colored according to ConSurf Conservation Grade (See documentation):
(Click to enlarge 🔍) | | Node1 | Node2 | LinkStrength | Comm | IsNode1Hub? | IsNode2Hub? | Node1Cons | Node2Cons | Node1Shell | Node2Shell | | L:L:?1 | R:R:L26 | 3.64 | 2 | Yes | No | 0 | 8 | 0 | 1 | | L:L:?1 | R:R:K76 | 12.06 | 2 | Yes | Yes | 0 | 8 | 0 | 1 | | L:L:?1 | R:R:S80 | 10.86 | 2 | Yes | No | 0 | 8 | 0 | 1 | | L:L:?1 | R:R:V83 | 7.84 | 2 | Yes | No | 0 | 9 | 0 | 1 | | L:L:?1 | R:R:Y87 | 6.94 | 2 | Yes | No | 0 | 8 | 0 | 1 | | L:L:?1 | R:R:M112 | 7.44 | 2 | Yes | Yes | 0 | 8 | 0 | 1 | | L:L:?1 | R:R:T181 | 4.85 | 2 | Yes | Yes | 0 | 9 | 0 | 1 | | L:L:?1 | R:R:W182 | 8.99 | 2 | Yes | Yes | 0 | 9 | 0 | 1 | | L:L:?1 | R:R:L306 | 4.55 | 2 | Yes | No | 0 | 7 | 0 | 1 | | L:L:?1 | R:R:R310 | 5.59 | 2 | Yes | No | 0 | 9 | 0 | 1 | | L:L:?1 | R:R:L312 | 4.55 | 2 | Yes | No | 0 | 8 | 0 | 1 | | L:L:?1 | R:R:S313 | 13.83 | 2 | Yes | No | 0 | 9 | 0 | 1 | | R:R:A20 | R:R:Y87 | 6.67 | 0 | No | No | 6 | 8 | 2 | 1 | | R:R:M22 | R:R:R310 | 11.17 | 0 | No | No | 3 | 9 | 2 | 1 | | R:R:F27 | R:R:S80 | 11.89 | 0 | No | No | 8 | 8 | 2 | 1 | | R:R:G30 | R:R:K76 | 3.49 | 2 | No | Yes | 9 | 8 | 2 | 1 | | R:R:G30 | R:R:I317 | 5.29 | 2 | No | No | 9 | 9 | 2 | 2 | | R:R:K76 | R:R:S316 | 4.59 | 2 | Yes | No | 8 | 8 | 1 | 2 | | R:R:I317 | R:R:K76 | 5.82 | 2 | No | Yes | 9 | 8 | 2 | 1 | | R:R:A86 | R:R:T181 | 3.36 | 0 | No | Yes | 6 | 9 | 2 | 1 | | R:R:T181 | R:R:Y87 | 14.98 | 2 | Yes | No | 9 | 8 | 1 | 1 | | R:R:F108 | R:R:M112 | 3.73 | 0 | Yes | Yes | 6 | 8 | 2 | 1 | | R:R:F184 | R:R:M112 | 6.22 | 2 | Yes | Yes | 8 | 8 | 2 | 1 | | R:R:L309 | R:R:M112 | 4.24 | 2 | Yes | Yes | 7 | 8 | 2 | 1 | | R:R:W182 | R:R:Y177 | 17.36 | 2 | Yes | Yes | 9 | 9 | 1 | 2 | | R:R:F184 | R:R:Y177 | 5.16 | 2 | Yes | Yes | 8 | 9 | 2 | 2 | | R:R:P179 | R:R:T181 | 5.25 | 0 | No | Yes | 8 | 9 | 2 | 1 | | R:R:L306 | R:R:W182 | 5.69 | 2 | No | Yes | 7 | 9 | 1 | 1 | | R:R:L309 | R:R:W182 | 7.97 | 2 | Yes | Yes | 7 | 9 | 2 | 1 | | R:R:F184 | R:R:L309 | 6.09 | 2 | Yes | Yes | 8 | 7 | 2 | 2 | | R:R:L306 | R:R:R310 | 12.15 | 2 | No | No | 7 | 9 | 1 | 1 | | R:R:L309 | R:R:V281 | 2.98 | 2 | Yes | No | 7 | 8 | 2 | 2 | | R:R:L312 | R:R:V281 | 2.98 | 0 | No | No | 8 | 8 | 1 | 2 | | L:L:?1 | R:R:G23 | 2.25 | 2 | Yes | No | 0 | 8 | 0 | 1 | | L:L:?1 | R:R:G116 | 2.25 | 2 | Yes | No | 0 | 8 | 0 | 1 | | R:R:G23 | R:R:G24 | 2.11 | 0 | No | No | 8 | 4 | 1 | 2 | | R:R:P81 | R:R:S80 | 1.78 | 0 | No | No | 8 | 8 | 2 | 1 | | R:R:T181 | R:R:V14 | 1.59 | 2 | Yes | No | 9 | 4 | 1 | 2 | |
| Statistics | Value |
|---|
| Average Number Of Links | 14.00 | | Average Number Of Links With An Hub | 4.00 | | Average Interaction Strength | 6.83 | | Average Nodes In Shell | 31.00 | | Average Hubs In Shell | 9.00 | | Average Links In Shell | 38.00 | | Average Links Mediated by Hubs In Shell | 30.00 |
| 
Physico-chemical properties of the nodes interacting with this ligand (click to enlarge 🔍) | 
Location of the nodes interacting with this ligand (click to enlarge 🔍) |
|


