- Press and hold down the left mouse button and move the mouse to rotate.
- Press and hold down the central mouse button and move the mouse to translate.
- Use the mouse wheel to zoom in or out.
- Click on any atom on your molecule and the corresponding residue label will be placed beneath the viewer.
| Color | ConSurf Grade |
| No Conservation data available | |
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 | |
| 6 | |
| 7 | |
| 8 | |
| 9 |
Index: link id, click on each number to highlight the corresponding link in the 3D visualization.
Node1 Node2: the two nodes of the corresponding link.
Int. Strength: the interaction strength between the two nodes.
Hub1?, Hub2?: "Yes" if the corresponding node has more than 3 links, otherwise "No".
Community: the id of the community the link belong to, otherwise 0.
ConSurf1, ConSurf2: these columns report the ConSurf conservation grades of the two nodes involved in a link.
| Index | Node1 | Node2 | Int. Strength | Hub1? | Hub2? | Community | ConSurf1 | ConSurf2 |
|---|---|---|---|---|---|---|---|---|
| 1 | L:L:S36 | R:R:S43 | 6.52 | No | No | 0 | 4 | 5 |
| 2 | L:L:V37 | R:R:G111 | 3.68 | No | No | 0 | 1 | 4 |
| 3 | L:L:V37 | R:R:S193 | 3.23 | No | No | 0 | 1 | 3 |
| 4 | L:L:E40 | L:L:T39 | 2.82 | Yes | No | 0 | 6 | 2 |
| 5 | L:L:E40 | L:L:R42 | 4.65 | Yes | Yes | 2 | 6 | 8 |
| 6 | L:L:E40 | R:R:R208 | 10.47 | Yes | No | 0 | 6 | 4 |
| 7 | L:L:E40 | R:R:R212 | 3.49 | Yes | Yes | 2 | 6 | 5 |
| 8 | L:L:E40 | R:R:D274 | 6.5 | Yes | Yes | 2 | 6 | 4 |
| 9 | L:L:E40 | R:R:R278 | 6.98 | Yes | Yes | 2 | 6 | 4 |
| 10 | L:L:H68 | L:L:L41 | 3.86 | Yes | No | 0 | 8 | 6 |
| 11 | L:L:L41 | R:R:Y197 | 19.93 | No | No | 0 | 6 | 3 |
| 12 | L:L:R42 | R:R:D274 | 8.34 | Yes | Yes | 2 | 8 | 4 |
| 13 | L:L:R42 | R:R:R278 | 17.06 | Yes | Yes | 2 | 8 | 4 |
| 14 | L:L:R42 | R:R:R289 | 10.66 | Yes | Yes | 2 | 8 | 3 |
| 15 | L:L:R42 | R:R:D293 | 5.96 | Yes | No | 0 | 8 | 4 |
| 16 | L:L:C43 | L:L:C69 | 7.28 | No | No | 7 | 9 | 9 |
| 17 | L:L:C43 | L:L:E73 | 9.12 | No | Yes | 7 | 9 | 9 |
| 18 | L:L:K63 | L:L:Q44 | 4.07 | Yes | Yes | 0 | 5 | 5 |
| 19 | L:L:E73 | L:L:Q44 | 3.82 | Yes | Yes | 0 | 9 | 5 |
| 20 | L:L:Q44 | R:R:N191 | 10.56 | Yes | No | 0 | 5 | 2 |
| 21 | L:L:C45 | L:L:E73 | 4.56 | No | Yes | 0 | 9 | 9 |
| 22 | L:L:C45 | L:L:C85 | 5.46 | No | No | 0 | 9 | 9 |
| 23 | L:L:L46 | L:L:Q47 | 6.65 | No | No | 0 | 5 | 4 |
| 24 | L:L:L46 | R:R:E284 | 3.98 | No | Yes | 2 | 5 | 3 |
| 25 | L:L:L46 | R:R:R289 | 3.64 | No | Yes | 2 | 5 | 3 |
| 26 | L:L:N87 | L:L:T48 | 2.92 | Yes | No | 0 | 7 | 8 |
| 27 | L:L:L49 | R:R:A36 | 4.73 | No | No | 0 | 5 | 5 |
| 28 | L:L:I52 | L:L:I57 | 8.83 | No | No | 0 | 8 | 8 |
| 29 | L:L:I52 | L:L:L78 | 2.85 | No | Yes | 0 | 8 | 7 |
| 30 | L:L:I52 | L:L:L86 | 4.28 | No | Yes | 0 | 8 | 9 |
| 31 | L:L:H53 | L:L:P54 | 7.63 | No | No | 0 | 4 | 5 |
| 32 | L:L:H53 | L:L:K55 | 5.24 | No | No | 0 | 4 | 5 |
| 33 | L:L:I96 | L:L:P54 | 3.39 | No | No | 9 | 5 | 5 |
| 34 | L:L:M100 | L:L:P54 | 3.35 | No | No | 9 | 5 | 5 |
| 35 | L:L:L78 | L:L:N56 | 9.61 | Yes | No | 0 | 7 | 4 |
| 36 | L:L:S59 | L:L:T77 | 4.8 | No | No | 0 | 4 | 8 |
| 37 | L:L:S59 | N:N:N61 | 2.98 | No | No | 0 | 4 | 6 |
| 38 | L:L:I97 | L:L:V60 | 3.07 | No | No | 0 | 7 | 6 |
| 39 | L:L:K63 | L:L:N61 | 4.2 | Yes | No | 1 | 5 | 6 |
| 40 | L:L:N61 | N:N:S59 | 2.98 | No | No | 1 | 6 | 4 |
| 41 | L:L:N61 | N:N:N61 | 12.26 | No | No | 0 | 6 | 6 |
| 42 | L:L:V62 | L:L:V74 | 6.41 | No | No | 0 | 7 | 8 |
| 43 | L:L:H68 | L:L:K63 | 5.24 | Yes | Yes | 1 | 8 | 5 |
| 44 | L:L:K63 | N:N:S59 | 3.06 | Yes | No | 1 | 5 | 4 |
| 45 | L:L:P65 | L:L:S64 | 3.56 | No | No | 0 | 6 | 5 |
| 46 | L:L:S64 | N:N:M100 | 4.6 | No | No | 0 | 5 | 5 |
| 47 | L:L:P65 | L:L:T72 | 6.99 | No | No | 0 | 6 | 5 |
| 48 | L:L:P65 | N:N:S103 | 7.13 | No | No | 0 | 6 | 4 |
| 49 | L:L:G66 | L:L:P67 | 4.06 | No | Yes | 0 | 8 | 6 |
| 50 | L:L:H68 | L:L:P67 | 4.58 | Yes | Yes | 1 | 8 | 6 |
| 51 | L:L:P67 | N:N:Q58 | 3.16 | Yes | No | 1 | 6 | 4 |
| 52 | L:L:P67 | R:R:R185 | 2.88 | Yes | No | 1 | 6 | 4 |
| 53 | L:L:P67 | R:R:V187 | 3.53 | Yes | No | 1 | 6 | 3 |
| 54 | L:L:P67 | R:R:D199 | 11.27 | Yes | No | 1 | 6 | 4 |
| 55 | L:L:H68 | N:N:Q58 | 3.71 | Yes | No | 1 | 8 | 4 |
| 56 | L:L:C69 | L:L:E73 | 9.12 | No | Yes | 7 | 9 | 9 |
| 57 | L:L:P88 | L:L:T72 | 5.25 | No | No | 0 | 9 | 5 |
| 58 | L:L:L86 | L:L:V74 | 4.47 | Yes | No | 0 | 9 | 8 |
| 59 | L:L:A76 | L:L:L86 | 3.15 | No | Yes | 0 | 8 | 9 |
| 60 | L:L:L78 | L:L:N80 | 6.87 | Yes | No | 8 | 7 | 5 |
| 61 | L:L:L78 | L:L:R82 | 3.64 | Yes | No | 8 | 7 | 4 |
| 62 | L:L:N80 | L:L:R82 | 3.62 | No | No | 8 | 5 | 4 |
| 63 | L:L:A84 | L:L:R82 | 4.15 | No | No | 0 | 6 | 4 |
| 64 | L:L:K83 | R:R:P38 | 3.35 | No | No | 0 | 4 | 7 |
| 65 | L:L:K83 | R:R:E40 | 5.4 | No | No | 0 | 4 | 4 |
| 66 | L:L:L86 | L:L:V93 | 8.94 | Yes | No | 0 | 9 | 8 |
| 67 | L:L:N87 | L:L:P88 | 6.52 | Yes | No | 0 | 7 | 9 |
| 68 | L:L:A89 | L:L:N87 | 4.69 | No | Yes | 0 | 4 | 7 |
| 69 | L:L:N87 | L:L:S90 | 4.47 | Yes | Yes | 0 | 7 | 8 |
| 70 | L:L:P91 | L:L:S90 | 3.56 | No | Yes | 0 | 5 | 8 |
| 71 | L:L:I92 | L:L:S90 | 4.64 | No | Yes | 0 | 4 | 8 |
| 72 | L:L:S90 | L:L:V93 | 4.85 | Yes | No | 0 | 8 | 8 |
| 73 | L:L:I92 | L:L:I96 | 2.94 | No | No | 0 | 4 | 5 |
| 74 | L:L:I97 | L:L:V93 | 3.07 | No | No | 0 | 7 | 8 |
| 75 | L:L:E98 | L:L:K94 | 4.05 | No | Yes | 0 | 3 | 6 |
| 76 | L:L:K94 | N:N:E98 | 10.8 | Yes | No | 10 | 6 | 3 |
| 77 | L:L:K94 | N:N:L101 | 7.05 | Yes | No | 0 | 6 | 6 |
| 78 | L:L:K94 | N:N:N102 | 4.2 | Yes | No | 10 | 6 | 4 |
| 79 | L:L:I96 | L:L:M100 | 2.92 | No | No | 9 | 5 | 5 |
| 80 | L:L:I97 | L:L:L101 | 2.85 | No | Yes | 0 | 7 | 6 |
| 81 | L:L:K99 | L:L:M100 | 2.88 | No | No | 0 | 5 | 5 |
| 82 | L:L:L101 | N:N:V62 | 2.98 | Yes | No | 0 | 6 | 7 |
| 83 | L:L:L101 | N:N:K94 | 8.46 | Yes | No | 0 | 6 | 6 |
| 84 | L:L:L101 | N:N:I97 | 5.71 | Yes | No | 0 | 6 | 7 |
| 85 | N:N:C43 | N:N:C69 | 7.28 | No | No | 11 | 9 | 9 |
| 86 | N:N:C43 | N:N:E73 | 6.08 | No | Yes | 11 | 9 | 9 |
| 87 | N:N:K63 | N:N:Q44 | 4.07 | No | No | 0 | 5 | 5 |
| 88 | N:N:C45 | N:N:E73 | 9.12 | No | Yes | 0 | 9 | 9 |
| 89 | N:N:C45 | N:N:I75 | 3.27 | No | No | 0 | 9 | 9 |
| 90 | N:N:C45 | N:N:C85 | 7.28 | No | No | 0 | 9 | 9 |
| 91 | N:N:N87 | N:N:T48 | 5.85 | Yes | No | 0 | 7 | 8 |
| 92 | N:N:L49 | N:N:Q50 | 9.32 | No | No | 0 | 5 | 7 |
| 93 | N:N:G51 | N:N:Q50 | 3.29 | No | No | 0 | 3 | 7 |
| 94 | N:N:I52 | N:N:I57 | 7.36 | Yes | No | 0 | 8 | 8 |
| 95 | N:N:I52 | N:N:L78 | 4.28 | Yes | No | 0 | 8 | 7 |
| 96 | N:N:I52 | N:N:L86 | 4.28 | Yes | Yes | 0 | 8 | 9 |
| 97 | N:N:H53 | N:N:P54 | 6.1 | No | No | 0 | 4 | 5 |
| 98 | N:N:H53 | N:N:K55 | 6.55 | No | No | 0 | 4 | 5 |
| 99 | N:N:H53 | N:N:N56 | 8.93 | No | No | 0 | 4 | 4 |
| 100 | N:N:I96 | N:N:P54 | 8.47 | No | No | 0 | 5 | 5 |
| 101 | N:N:A76 | N:N:I57 | 4.87 | No | No | 0 | 8 | 8 |
| 102 | N:N:K79 | N:N:Q58 | 5.42 | No | No | 0 | 8 | 4 |
| 103 | N:N:S59 | N:N:T77 | 3.2 | No | No | 0 | 4 | 8 |
| 104 | N:N:I97 | N:N:V60 | 3.07 | No | No | 0 | 7 | 6 |
| 105 | N:N:M100 | N:N:V60 | 4.56 | No | No | 0 | 5 | 6 |
| 106 | N:N:V62 | N:N:V74 | 6.41 | No | No | 0 | 7 | 8 |
| 107 | N:N:E73 | N:N:K63 | 4.05 | Yes | No | 0 | 9 | 5 |
| 108 | N:N:P65 | N:N:S64 | 3.56 | No | No | 0 | 6 | 5 |
| 109 | N:N:P65 | N:N:T72 | 5.25 | No | No | 0 | 6 | 5 |
| 110 | N:N:G66 | N:N:P67 | 4.06 | No | No | 0 | 8 | 6 |
| 111 | N:N:C69 | N:N:E73 | 4.56 | No | Yes | 11 | 9 | 9 |
| 112 | N:N:L86 | N:N:V74 | 2.98 | Yes | No | 0 | 9 | 8 |
| 113 | N:N:A76 | N:N:L86 | 3.15 | No | Yes | 0 | 8 | 9 |
| 114 | N:N:K83 | N:N:T77 | 9.01 | No | No | 0 | 4 | 8 |
| 115 | N:N:L78 | N:N:N80 | 8.24 | No | No | 0 | 7 | 5 |
| 116 | N:N:G81 | R:R:Y188 | 5.79 | No | No | 0 | 7 | 1 |
| 117 | N:N:L86 | N:N:V93 | 8.94 | Yes | No | 0 | 9 | 8 |
| 118 | N:N:N87 | N:N:P88 | 6.52 | Yes | No | 0 | 7 | 9 |
| 119 | N:N:A89 | N:N:N87 | 4.69 | No | Yes | 0 | 4 | 7 |
| 120 | N:N:N87 | N:N:S90 | 4.47 | Yes | Yes | 0 | 7 | 8 |
| 121 | N:N:P91 | N:N:S90 | 3.56 | No | Yes | 0 | 5 | 8 |
| 122 | N:N:I92 | N:N:S90 | 4.64 | No | Yes | 0 | 4 | 8 |
| 123 | N:N:S90 | N:N:V93 | 6.46 | Yes | No | 0 | 8 | 8 |
| 124 | N:N:I92 | N:N:I96 | 2.94 | No | No | 0 | 4 | 5 |
| 125 | N:N:E98 | N:N:N102 | 3.94 | No | No | 10 | 3 | 4 |
| 126 | R:R:C286 | R:R:C39 | 7.28 | No | No | 0 | 6 | 8 |
| 127 | R:R:C39 | R:R:R289 | 11.14 | No | Yes | 0 | 8 | 3 |
| 128 | R:R:E45 | R:R:K48 | 4.05 | No | No | 0 | 4 | 4 |
| 129 | R:R:I301 | R:R:V51 | 7.68 | No | No | 0 | 5 | 5 |
| 130 | R:R:A105 | R:R:V52 | 3.39 | No | No | 0 | 7 | 5 |
| 131 | R:R:I301 | R:R:I54 | 2.94 | No | No | 0 | 5 | 5 |
| 132 | R:R:L101 | R:R:Y55 | 5.86 | No | Yes | 0 | 8 | 7 |
| 133 | R:R:A105 | R:R:Y55 | 4 | No | Yes | 0 | 7 | 7 |
| 134 | R:R:I301 | R:R:Y55 | 4.84 | No | Yes | 0 | 5 | 7 |
| 135 | R:R:I304 | R:R:Y55 | 8.46 | No | Yes | 0 | 7 | 7 |
| 136 | R:R:L57 | R:R:L60 | 4.15 | No | No | 0 | 5 | 4 |
| 137 | R:R:L101 | R:R:V58 | 2.98 | No | No | 0 | 8 | 7 |
| 138 | R:R:I304 | R:R:V58 | 6.14 | No | No | 0 | 7 | 7 |
| 139 | R:R:C308 | R:R:V58 | 5.12 | No | No | 0 | 7 | 7 |
| 140 | R:R:F59 | R:R:L63 | 6.09 | No | No | 0 | 7 | 5 |
| 141 | R:R:F59 | R:R:P102 | 10.11 | No | No | 0 | 7 | 9 |
| 142 | R:R:C308 | R:R:L61 | 3.17 | No | No | 0 | 7 | 5 |
| 143 | R:R:S307 | R:R:S62 | 6.52 | No | No | 0 | 8 | 8 |
| 144 | R:R:C308 | R:R:S62 | 5.16 | No | No | 0 | 7 | 8 |
| 145 | R:R:L63 | R:R:L95 | 2.77 | No | No | 0 | 5 | 7 |
| 146 | R:R:L64 | R:R:S67 | 3 | No | No | 0 | 4 | 3 |
| 147 | R:R:L64 | R:R:L68 | 6.92 | No | No | 0 | 4 | 6 |
| 148 | R:R:G65 | R:R:L68 | 3.42 | No | No | 0 | 7 | 6 |
| 149 | R:R:G65 | R:R:P311 | 4.06 | No | No | 0 | 7 | 9 |
| 150 | R:R:A91 | R:R:N66 | 3.13 | No | No | 0 | 8 | 9 |
| 151 | R:R:D94 | R:R:N66 | 6.73 | Yes | No | 0 | 9 | 9 |
| 152 | R:R:L95 | R:R:N66 | 9.61 | No | No | 0 | 7 | 9 |
| 153 | R:R:F316 | R:R:L68 | 7.31 | Yes | No | 0 | 6 | 6 |
| 154 | R:R:A91 | R:R:V69 | 3.39 | No | No | 0 | 8 | 8 |
| 155 | R:R:F316 | R:R:V69 | 3.93 | Yes | No | 0 | 6 | 8 |
| 156 | R:R:L95 | R:R:M70 | 2.83 | No | No | 0 | 7 | 6 |
| 157 | R:R:F321 | R:R:V72 | 5.24 | No | No | 0 | 8 | 7 |
| 158 | R:R:I73 | R:R:L88 | 11.42 | No | No | 0 | 6 | 5 |
| 159 | R:R:F321 | R:R:I73 | 7.54 | No | No | 0 | 8 | 6 |
| 160 | R:R:L74 | R:R:R80 | 3.64 | No | No | 0 | 2 | 7 |
| 161 | R:R:L74 | R:R:L88 | 4.15 | No | No | 0 | 2 | 5 |
| 162 | R:R:I328 | R:R:Y75 | 4.84 | No | No | 0 | 5 | 4 |
| 163 | R:R:R77 | R:R:R80 | 3.2 | No | No | 0 | 6 | 7 |
| 164 | R:R:D84 | R:R:S81 | 8.83 | No | No | 0 | 8 | 7 |
| 165 | R:R:D143 | R:R:V82 | 5.84 | No | No | 0 | 8 | 7 |
| 166 | R:R:V162 | R:R:V82 | 9.62 | No | No | 0 | 7 | 7 |
| 167 | R:R:N89 | R:R:V85 | 4.43 | No | No | 0 | 9 | 6 |
| 168 | R:R:V162 | R:R:V85 | 3.21 | No | No | 0 | 7 | 6 |
| 169 | R:R:C139 | R:R:Y86 | 4.03 | No | Yes | 0 | 5 | 7 |
| 170 | R:R:I140 | R:R:Y86 | 6.04 | No | Yes | 0 | 9 | 7 |
| 171 | R:R:D143 | R:R:Y86 | 4.6 | No | Yes | 0 | 8 | 7 |
| 172 | R:R:V162 | R:R:Y86 | 8.83 | No | Yes | 0 | 7 | 7 |
| 173 | R:R:A315 | R:R:L87 | 3.15 | No | No | 0 | 7 | 7 |
| 174 | R:R:C166 | R:R:N89 | 7.87 | No | No | 0 | 7 | 9 |
| 175 | R:R:D94 | R:R:L90 | 6.79 | Yes | Yes | 12 | 9 | 9 |
| 176 | R:R:L136 | R:R:L90 | 2.77 | No | Yes | 0 | 8 | 9 |
| 177 | R:R:L137 | R:R:L90 | 5.54 | No | Yes | 0 | 8 | 9 |
| 178 | R:R:L90 | R:R:N310 | 5.49 | Yes | No | 12 | 9 | 9 |
| 179 | R:R:D94 | R:R:S307 | 4.42 | Yes | No | 0 | 9 | 8 |
| 180 | R:R:D94 | R:R:N310 | 12.12 | Yes | No | 12 | 9 | 9 |
| 181 | R:R:L96 | R:R:N129 | 6.87 | No | Yes | 0 | 7 | 8 |
| 182 | R:R:F97 | R:R:K126 | 7.44 | No | No | 0 | 6 | 6 |
| 183 | R:R:F97 | R:R:H306 | 9.05 | No | No | 0 | 6 | 9 |
| 184 | R:R:A98 | R:R:L101 | 3.15 | No | No | 0 | 5 | 8 |
| 185 | R:R:T100 | R:R:W104 | 3.64 | Yes | No | 13 | 7 | 6 |
| 186 | R:R:T100 | R:R:V122 | 4.76 | Yes | No | 0 | 7 | 5 |
| 187 | R:R:K126 | R:R:T100 | 4.5 | No | Yes | 13 | 6 | 7 |
| 188 | R:R:K126 | R:R:W104 | 17.4 | No | No | 13 | 6 | 6 |
| 189 | R:R:A106 | R:R:F114 | 2.77 | No | Yes | 0 | 4 | 6 |
| 190 | R:R:S107 | R:R:W112 | 7.41 | No | Yes | 0 | 5 | 9 |
| 191 | R:R:D297 | R:R:K108 | 2.77 | No | No | 0 | 4 | 5 |
| 192 | R:R:N110 | R:R:V109 | 8.87 | No | No | 0 | 4 | 4 |
| 193 | R:R:F114 | R:R:W112 | 5.01 | Yes | Yes | 0 | 6 | 9 |
| 194 | R:R:G115 | R:R:W112 | 2.81 | No | Yes | 0 | 8 | 9 |
| 195 | R:R:C119 | R:R:W112 | 7.84 | No | Yes | 5 | 9 | 9 |
| 196 | R:R:T186 | R:R:W112 | 4.85 | Yes | Yes | 5 | 4 | 9 |
| 197 | R:R:C196 | R:R:W112 | 7.84 | No | Yes | 5 | 9 | 9 |
| 198 | R:R:F114 | R:R:L118 | 3.65 | Yes | No | 0 | 6 | 6 |
| 199 | R:R:T116 | R:R:T186 | 4.71 | No | Yes | 0 | 3 | 4 |
| 200 | R:R:F117 | R:R:V121 | 5.24 | No | No | 0 | 1 | 5 |
| 201 | R:R:C119 | R:R:T186 | 3.38 | No | Yes | 5 | 9 | 4 |
| 202 | R:R:C119 | R:R:C196 | 7.28 | No | No | 5 | 9 | 9 |
| 203 | R:R:R184 | R:R:S123 | 10.54 | Yes | No | 0 | 5 | 5 |
| 204 | R:R:L124 | R:R:L181 | 16.61 | No | No | 0 | 6 | 5 |
| 205 | R:R:L125 | R:R:N129 | 5.49 | No | Yes | 0 | 4 | 8 |
| 206 | R:R:E127 | R:R:Y131 | 8.98 | No | Yes | 0 | 4 | 7 |
| 207 | R:R:A177 | R:R:E127 | 3.02 | No | No | 0 | 7 | 4 |
| 208 | R:R:E127 | R:R:V180 | 12.84 | No | No | 0 | 4 | 4 |
| 209 | R:R:N129 | R:R:V128 | 4.43 | Yes | No | 0 | 8 | 5 |
| 210 | R:R:S173 | R:R:V128 | 3.23 | No | No | 0 | 7 | 5 |
| 211 | R:R:F130 | R:R:Y131 | 9.28 | Yes | Yes | 0 | 7 | 7 |
| 212 | R:R:F130 | R:R:I134 | 6.28 | Yes | Yes | 3 | 7 | 8 |
| 213 | R:R:F130 | R:R:W264 | 10.02 | Yes | Yes | 3 | 7 | 8 |
| 214 | R:R:L176 | R:R:Y131 | 12.89 | No | Yes | 0 | 4 | 7 |
| 215 | R:R:P215 | R:R:Y131 | 4.17 | No | Yes | 0 | 5 | 7 |
| 216 | R:R:S132 | R:R:W170 | 14.83 | No | No | 0 | 6 | 9 |
| 217 | R:R:S132 | R:R:S173 | 6.52 | No | No | 0 | 6 | 7 |
| 218 | R:R:I134 | R:R:P223 | 3.39 | Yes | No | 0 | 8 | 9 |
| 219 | R:R:F260 | R:R:I134 | 3.77 | Yes | Yes | 3 | 9 | 8 |
| 220 | R:R:I134 | R:R:W264 | 5.87 | Yes | Yes | 3 | 8 | 8 |
| 221 | R:R:I169 | R:R:L135 | 5.71 | No | No | 0 | 6 | 6 |
| 222 | R:R:I169 | R:R:L136 | 2.85 | No | No | 0 | 6 | 8 |
| 223 | R:R:L137 | R:R:M227 | 5.65 | No | No | 0 | 8 | 8 |
| 224 | R:R:L137 | R:R:Y314 | 9.38 | No | Yes | 0 | 8 | 9 |
| 225 | R:R:C139 | R:R:I169 | 3.27 | No | No | 0 | 5 | 6 |
| 226 | R:R:I140 | R:R:R144 | 6.26 | No | No | 4 | 9 | 9 |
| 227 | R:R:I140 | R:R:Y314 | 6.04 | No | Yes | 4 | 9 | 9 |
| 228 | R:R:M227 | R:R:S141 | 4.6 | No | No | 0 | 8 | 9 |
| 229 | R:R:C230 | R:R:S141 | 3.44 | No | No | 0 | 7 | 9 |
| 230 | R:R:S141 | R:R:Y231 | 13.99 | No | Yes | 0 | 9 | 9 |
| 231 | R:R:C230 | R:R:V142 | 3.42 | No | No | 0 | 7 | 6 |
| 232 | R:R:D143 | R:R:R159 | 9.53 | No | No | 0 | 8 | 3 |
| 233 | R:R:R144 | R:R:Y231 | 9.26 | No | Yes | 0 | 9 | 9 |
| 234 | R:R:R144 | R:R:Y314 | 6.17 | No | Yes | 4 | 9 | 9 |
| 235 | R:R:L146 | R:R:Y145 | 4.69 | No | Yes | 0 | 5 | 8 |
| 236 | R:R:V149 | R:R:Y145 | 7.57 | Yes | Yes | 6 | 7 | 8 |
| 237 | R:R:H150 | R:R:Y145 | 9.8 | Yes | Yes | 6 | 6 | 8 |
| 238 | R:R:C230 | R:R:Y145 | 6.72 | No | Yes | 0 | 7 | 8 |
| 239 | R:R:F233 | R:R:Y145 | 3.09 | No | Yes | 0 | 4 | 8 |
| 240 | R:R:T234 | R:R:Y145 | 3.75 | Yes | Yes | 6 | 8 | 8 |
| 241 | R:R:L146 | R:R:L155 | 2.77 | No | No | 0 | 5 | 4 |
| 242 | R:R:A147 | R:R:R159 | 6.91 | No | No | 0 | 8 | 3 |
| 243 | R:R:I148 | R:R:T234 | 4.56 | No | Yes | 0 | 8 | 8 |
| 244 | R:R:I148 | R:R:L238 | 2.85 | No | No | 0 | 8 | 8 |
| 245 | R:R:H150 | R:R:V149 | 5.54 | Yes | Yes | 6 | 6 | 7 |
| 246 | R:R:T234 | R:R:V149 | 3.17 | Yes | Yes | 6 | 8 | 7 |
| 247 | R:R:T237 | R:R:V149 | 6.35 | No | Yes | 0 | 5 | 7 |
| 248 | R:R:H150 | R:R:R153 | 5.64 | Yes | No | 0 | 6 | 5 |
| 249 | R:R:H150 | R:R:L155 | 5.14 | Yes | No | 0 | 6 | 4 |
| 250 | R:R:Q157 | R:R:T154 | 2.83 | No | No | 0 | 7 | 5 |
| 251 | R:R:K158 | R:R:L155 | 4.23 | No | No | 0 | 6 | 4 |
| 252 | R:R:L161 | R:R:Y160 | 5.86 | No | No | 0 | 4 | 5 |
| 253 | R:R:L172 | R:R:L176 | 6.92 | No | No | 0 | 4 | 4 |
| 254 | R:R:L211 | R:R:P179 | 8.21 | No | No | 0 | 4 | 8 |
| 255 | R:R:R184 | R:R:V180 | 6.54 | Yes | No | 0 | 5 | 4 |
| 256 | R:R:F183 | R:R:L182 | 4.87 | No | No | 0 | 5 | 4 |
| 257 | R:R:F183 | R:R:M200 | 13.68 | No | No | 0 | 5 | 5 |
| 258 | R:R:C196 | R:R:R184 | 2.79 | No | Yes | 0 | 9 | 5 |
| 259 | R:R:E198 | R:R:R184 | 19.77 | No | Yes | 0 | 5 | 5 |
| 260 | R:R:R185 | R:R:V187 | 3.92 | No | No | 1 | 4 | 3 |
| 261 | R:R:D199 | R:R:R185 | 19.06 | No | No | 1 | 4 | 4 |
| 262 | R:R:T186 | R:R:Y188 | 3.75 | Yes | No | 0 | 4 | 1 |
| 263 | R:R:P194 | R:R:Y188 | 11.13 | No | No | 0 | 5 | 1 |
| 264 | R:R:N191 | R:R:S190 | 5.96 | No | No | 0 | 2 | 2 |
| 265 | R:R:P194 | R:R:S193 | 3.56 | No | No | 0 | 5 | 3 |
| 266 | R:R:R208 | R:R:Y197 | 18.52 | No | No | 0 | 4 | 3 |
| 267 | R:R:R278 | R:R:Y197 | 3.09 | Yes | No | 0 | 4 | 3 |
| 268 | R:R:E198 | R:R:R208 | 10.47 | No | No | 0 | 5 | 4 |
| 269 | R:R:M200 | R:R:T204 | 3.01 | No | No | 0 | 5 | 3 |
| 270 | R:R:M200 | R:R:W207 | 6.98 | No | No | 0 | 5 | 3 |
| 271 | R:R:N203 | R:R:N206 | 10.9 | No | No | 0 | 4 | 1 |
| 272 | R:R:L210 | R:R:W207 | 3.42 | No | No | 0 | 2 | 3 |
| 273 | R:R:M209 | R:R:T275 | 7.53 | No | No | 0 | 3 | 4 |
| 274 | R:R:M209 | R:R:R278 | 3.72 | No | Yes | 0 | 3 | 4 |
| 275 | R:R:L271 | R:R:R212 | 10.93 | Yes | Yes | 0 | 6 | 5 |
| 276 | R:R:D274 | R:R:R212 | 10.72 | Yes | Yes | 2 | 4 | 5 |
| 277 | R:R:R212 | R:R:T275 | 6.47 | Yes | No | 0 | 5 | 4 |
| 278 | R:R:I213 | R:R:S217 | 6.19 | No | No | 0 | 4 | 4 |
| 279 | R:R:N268 | R:R:Q216 | 7.92 | No | No | 0 | 8 | 5 |
| 280 | R:R:L271 | R:R:Q216 | 5.32 | Yes | No | 0 | 6 | 5 |
| 281 | R:R:F220 | R:R:L224 | 3.65 | Yes | Yes | 3 | 8 | 7 |
| 282 | R:R:F220 | R:R:F260 | 4.29 | Yes | Yes | 3 | 8 | 9 |
| 283 | R:R:F220 | R:R:L265 | 3.65 | Yes | No | 0 | 8 | 6 |
| 284 | R:R:F220 | R:R:N268 | 15.71 | Yes | No | 0 | 8 | 8 |
| 285 | R:R:I221 | R:R:L225 | 2.85 | No | No | 0 | 4 | 3 |
| 286 | R:R:P223 | R:R:V222 | 3.53 | No | No | 0 | 9 | 5 |
| 287 | R:R:I226 | R:R:V222 | 3.07 | No | No | 0 | 6 | 5 |
| 288 | R:R:L224 | R:R:L228 | 4.15 | Yes | No | 0 | 7 | 4 |
| 289 | R:R:F260 | R:R:L224 | 4.87 | Yes | Yes | 3 | 9 | 7 |
| 290 | R:R:L224 | R:R:L261 | 4.15 | Yes | No | 0 | 7 | 6 |
| 291 | R:R:F260 | R:R:M227 | 11.2 | Yes | No | 0 | 9 | 8 |
| 292 | R:R:F229 | R:R:F233 | 7.5 | No | No | 0 | 4 | 4 |
| 293 | R:R:I253 | R:R:Y231 | 9.67 | No | Yes | 0 | 7 | 9 |
| 294 | R:R:V256 | R:R:Y231 | 3.79 | Yes | Yes | 0 | 8 | 9 |
| 295 | R:R:V257 | R:R:Y231 | 5.05 | No | Yes | 0 | 7 | 9 |
| 296 | R:R:I253 | R:R:T234 | 3.04 | No | Yes | 0 | 7 | 8 |
| 297 | R:R:L235 | R:R:M250 | 2.83 | No | Yes | 0 | 4 | 5 |
| 298 | R:R:I253 | R:R:L235 | 7.14 | No | No | 0 | 7 | 4 |
| 299 | R:R:F254 | R:R:L235 | 4.87 | No | No | 0 | 4 | 4 |
| 300 | R:R:L238 | R:R:M250 | 2.83 | No | Yes | 0 | 8 | 5 |
| 301 | R:R:F239 | R:R:M250 | 6.22 | No | Yes | 0 | 5 | 5 |
| 302 | R:R:G244 | R:R:Q245 | 3.29 | No | No | 0 | 4 | 4 |
| 303 | R:R:H247 | R:R:K246 | 7.86 | No | No | 0 | 5 | 7 |
| 304 | R:R:K246 | R:R:M250 | 5.76 | No | Yes | 0 | 7 | 5 |
| 305 | R:R:R248 | R:R:R322 | 3.2 | No | No | 0 | 7 | 8 |
| 306 | R:R:I317 | R:R:V252 | 7.68 | No | No | 0 | 6 | 7 |
| 307 | R:R:I313 | R:R:V256 | 3.07 | No | Yes | 4 | 7 | 8 |
| 308 | R:R:V256 | R:R:Y314 | 6.31 | Yes | Yes | 4 | 8 | 9 |
| 309 | R:R:I259 | R:R:L309 | 4.28 | No | No | 0 | 6 | 6 |
| 310 | R:R:I259 | R:R:I313 | 7.36 | No | No | 0 | 6 | 7 |
| 311 | R:R:F260 | R:R:W264 | 3.01 | Yes | Yes | 3 | 9 | 8 |
| 312 | R:R:W264 | R:R:Y267 | 2.89 | Yes | Yes | 0 | 8 | 7 |
| 313 | R:R:H306 | R:R:W264 | 7.41 | No | Yes | 0 | 9 | 8 |
| 314 | R:R:L265 | R:R:P266 | 3.28 | No | No | 0 | 6 | 9 |
| 315 | R:R:L265 | R:R:L269 | 5.54 | No | No | 0 | 6 | 6 |
| 316 | R:R:L302 | R:R:P266 | 3.28 | No | No | 0 | 7 | 9 |
| 317 | R:R:L271 | R:R:Y267 | 7.03 | Yes | Yes | 0 | 6 | 7 |
| 318 | R:R:T299 | R:R:Y267 | 4.99 | No | Yes | 0 | 6 | 7 |
| 319 | R:R:E300 | R:R:Y267 | 3.37 | No | Yes | 0 | 5 | 7 |
| 320 | R:R:A295 | R:R:V270 | 3.39 | No | No | 0 | 5 | 4 |
| 321 | R:R:L296 | R:R:V270 | 4.47 | No | No | 0 | 5 | 4 |
| 322 | R:R:T299 | R:R:V270 | 7.93 | No | No | 0 | 6 | 4 |
| 323 | R:R:L271 | R:R:L296 | 4.15 | Yes | No | 0 | 6 | 5 |
| 324 | R:R:L272 | R:R:L276 | 2.77 | No | No | 0 | 5 | 4 |
| 325 | R:R:D274 | R:R:R278 | 8.34 | Yes | Yes | 2 | 4 | 4 |
| 326 | R:R:L276 | R:R:V281 | 4.47 | No | No | 0 | 4 | 4 |
| 327 | R:R:E284 | R:R:M277 | 5.41 | Yes | Yes | 0 | 3 | 4 |
| 328 | R:R:M277 | R:R:R288 | 3.72 | Yes | No | 0 | 4 | 1 |
| 329 | R:R:I292 | R:R:M277 | 4.37 | No | Yes | 0 | 5 | 4 |
| 330 | R:R:T279 | R:R:V281 | 3.17 | No | No | 0 | 1 | 4 |
| 331 | R:R:E284 | R:R:Q280 | 11.47 | Yes | No | 0 | 3 | 2 |
| 332 | R:R:Q283 | R:R:T285 | 4.25 | No | No | 0 | 2 | 5 |
| 333 | R:R:E284 | R:R:R289 | 5.82 | Yes | Yes | 2 | 3 | 3 |
| 334 | R:R:R288 | R:R:T285 | 7.76 | No | No | 0 | 1 | 5 |
| 335 | R:R:E287 | R:R:H291 | 3.69 | No | No | 0 | 4 | 3 |
| 336 | R:R:H291 | R:R:N290 | 3.83 | No | No | 0 | 3 | 3 |
| 337 | R:R:E300 | R:R:I304 | 4.1 | No | No | 0 | 5 | 7 |
| 338 | R:R:H306 | R:R:S307 | 2.79 | No | No | 0 | 9 | 8 |
| 339 | R:R:N310 | R:R:P311 | 3.26 | No | No | 0 | 9 | 9 |
| 340 | R:R:F316 | R:R:L312 | 3.65 | Yes | No | 0 | 6 | 6 |
| 341 | R:R:I313 | R:R:Y314 | 3.63 | No | Yes | 4 | 7 | 9 |
| 342 | R:R:A315 | R:R:F316 | 2.77 | No | Yes | 0 | 7 | 6 |
| 343 | R:R:A315 | R:R:F321 | 8.32 | No | No | 0 | 7 | 8 |
| 344 | R:R:L325 | R:R:L326 | 4.15 | No | No | 0 | 6 | 4 |
| 345 | R:R:I328 | R:R:K327 | 2.91 | No | No | 0 | 5 | 4 |
| 346 | R:R:L44 | R:R:N47 | 2.75 | No | No | 0 | 2 | 4 |
| 347 | L:L:I75 | L:L:Q44 | 2.74 | No | Yes | 0 | 9 | 5 |
| 348 | R:R:E300 | R:R:K108 | 2.7 | No | No | 0 | 5 | 5 |
| 349 | R:R:H323 | R:R:K327 | 2.62 | No | No | 0 | 4 | 4 |
| 350 | R:R:C263 | R:R:W264 | 2.61 | No | Yes | 0 | 7 | 8 |
| 351 | R:R:A93 | R:R:W170 | 2.59 | No | No | 0 | 8 | 9 |
| 352 | R:R:R159 | R:R:T156 | 2.59 | No | No | 0 | 3 | 2 |
| 353 | N:N:E73 | N:N:Q71 | 2.55 | Yes | No | 0 | 9 | 5 |
| 354 | R:R:F50 | R:R:I46 | 2.51 | No | No | 0 | 5 | 5 |
| 355 | R:R:F114 | R:R:I103 | 2.51 | Yes | No | 0 | 6 | 6 |
| 356 | R:R:L44 | R:R:R294 | 2.43 | No | No | 0 | 2 | 2 |
| 357 | R:R:I165 | R:R:Y86 | 2.42 | No | Yes | 0 | 5 | 7 |
| 358 | R:R:F97 | R:R:N129 | 2.42 | No | Yes | 0 | 6 | 8 |
| 359 | N:N:N80 | N:N:R82 | 2.41 | No | No | 0 | 5 | 4 |
| 360 | R:R:N203 | R:R:W207 | 2.26 | No | No | 0 | 4 | 3 |
| 361 | R:R:H247 | R:R:R251 | 2.26 | No | No | 0 | 5 | 5 |
| 362 | R:R:A37 | R:R:P38 | 1.87 | No | No | 0 | 7 | 7 |
| 363 | R:R:A138 | R:R:P223 | 1.87 | No | No | 0 | 7 | 9 |
| 364 | R:R:G133 | R:R:S132 | 1.86 | No | No | 0 | 9 | 6 |
| 365 | R:R:G324 | R:R:V72 | 1.84 | No | No | 0 | 5 | 7 |
| 366 | R:R:G201 | R:R:T204 | 1.82 | No | No | 0 | 4 | 3 |
| 367 | L:L:C85 | R:R:A36 | 1.81 | No | No | 0 | 9 | 5 |
| 368 | L:L:A70 | R:R:A205 | 1.79 | No | No | 0 | 3 | 1 |
| 369 | R:R:P41 | R:R:V192 | 1.77 | No | No | 0 | 4 | 5 |
| 370 | L:L:G51 | L:L:I92 | 1.76 | No | No | 0 | 3 | 4 |
| 371 | R:R:G318 | R:R:I317 | 1.76 | No | No | 0 | 7 | 6 |
| 372 | L:L:P67 | R:R:T204 | 1.75 | Yes | No | 0 | 6 | 3 |
| 373 | N:N:P88 | N:N:T72 | 1.75 | No | No | 0 | 9 | 5 |
| 374 | R:R:G244 | R:R:M243 | 1.75 | No | No | 0 | 4 | 4 |
| 375 | L:L:A35 | R:R:S193 | 1.71 | No | No | 0 | 3 | 3 |
| 376 | R:R:G171 | R:R:L174 | 1.71 | No | No | 0 | 4 | 5 |
| 377 | R:R:G303 | R:R:L302 | 1.71 | No | No | 0 | 7 | 7 |
| 378 | L:L:A38 | R:R:V192 | 1.7 | No | No | 0 | 2 | 5 |
| 379 | L:L:G81 | L:L:N80 | 1.7 | No | No | 0 | 7 | 5 |
| 380 | N:N:G81 | N:N:N80 | 1.7 | No | No | 0 | 7 | 5 |
| 381 | L:L:I75 | R:R:P38 | 1.69 | No | No | 0 | 9 | 7 |
| 382 | N:N:C85 | N:N:T48 | 1.69 | No | No | 0 | 9 | 8 |
| 383 | L:L:P88 | N:N:L101 | 1.64 | No | No | 0 | 9 | 6 |
| 384 | R:R:L178 | R:R:P179 | 1.64 | No | No | 0 | 3 | 8 |
| 385 | R:R:L214 | R:R:P215 | 1.64 | No | No | 0 | 3 | 5 |
| 386 | R:R:G318 | R:R:Q319 | 1.64 | No | No | 0 | 7 | 5 |
| 387 | R:R:N191 | R:R:P41 | 1.63 | No | No | 0 | 2 | 4 |
| 388 | R:R:S189 | R:R:S190 | 1.63 | No | No | 0 | 1 | 2 |
| 389 | R:R:S76 | R:R:V78 | 1.62 | No | No | 0 | 4 | 5 |
| 390 | R:R:A273 | R:R:I292 | 1.62 | No | No | 0 | 5 | 5 |
| 391 | R:R:A241 | R:R:M243 | 1.61 | No | No | 0 | 3 | 4 |
| 392 | R:R:V109 | R:R:V52 | 1.6 | No | No | 0 | 4 | 5 |
| 393 | R:R:V252 | R:R:V256 | 1.6 | No | Yes | 0 | 7 | 8 |
| 394 | R:R:C263 | R:R:L305 | 1.59 | No | No | 0 | 7 | 5 |
| 395 | R:R:G324 | R:R:H323 | 1.59 | No | No | 0 | 5 | 4 |
| 396 | L:L:Q58 | N:N:P67 | 1.58 | No | No | 0 | 4 | 6 |
| 397 | R:R:A177 | R:R:L124 | 1.58 | No | No | 0 | 7 | 6 |
| 398 | R:R:A249 | R:R:L238 | 1.58 | No | No | 0 | 7 | 8 |
| 399 | R:R:T275 | R:R:T279 | 1.57 | No | No | 0 | 4 | 1 |
| 400 | L:L:A70 | R:R:N203 | 1.56 | No | No | 0 | 3 | 4 |
| 401 | R:R:A205 | R:R:N206 | 1.56 | No | No | 0 | 1 | 1 |
| 402 | R:R:I165 | R:R:S168 | 1.55 | No | No | 0 | 5 | 2 |
| 403 | R:R:K320 | R:R:S81 | 1.53 | No | No | 0 | 6 | 7 |
| 404 | R:R:F130 | R:R:G219 | 1.51 | Yes | No | 0 | 7 | 5 |
| 405 | R:R:K120 | R:R:T116 | 1.5 | No | No | 0 | 7 | 3 |
| 406 | R:R:G318 | R:R:R322 | 1.5 | No | No | 0 | 7 | 8 |
| 407 | R:R:L325 | R:R:V72 | 1.49 | No | No | 0 | 6 | 7 |
| 408 | R:R:L174 | R:R:V128 | 1.49 | No | No | 0 | 5 | 5 |
| 409 | R:R:L87 | R:R:T83 | 1.47 | No | No | 0 | 7 | 7 |
| 410 | R:R:L99 | R:R:T100 | 1.47 | No | Yes | 0 | 4 | 7 |
| 411 | R:R:I282 | R:R:M277 | 1.46 | No | Yes | 0 | 5 | 4 |
| 412 | L:L:Q58 | N:N:S64 | 1.44 | No | No | 0 | 4 | 5 |
| 413 | R:R:Q216 | R:R:S217 | 1.44 | No | No | 0 | 5 | 4 |
| 414 | R:R:K246 | R:R:M243 | 1.44 | No | No | 0 | 7 | 4 |
| 415 | N:N:I52 | N:N:L49 | 1.43 | Yes | No | 0 | 8 | 5 |
| 416 | R:R:Q157 | R:R:T156 | 1.42 | No | No | 0 | 7 | 2 |
| 417 | R:R:E287 | R:R:T285 | 1.41 | No | No | 0 | 4 | 5 |
| 418 | L:L:K79 | L:L:N56 | 1.4 | No | No | 0 | 8 | 4 |
| 419 | R:R:D84 | R:R:I73 | 1.4 | No | No | 0 | 8 | 6 |
| 420 | R:R:L92 | R:R:L96 | 1.38 | No | No | 0 | 6 | 7 |
| 421 | R:R:L174 | R:R:L175 | 1.38 | No | No | 0 | 5 | 4 |
| 422 | R:R:L326 | R:R:L329 | 1.38 | No | No | 0 | 4 | 6 |
| 423 | R:R:F218 | R:R:S217 | 1.32 | No | No | 0 | 4 | 4 |
| 424 | R:R:E42 | R:R:N290 | 1.31 | No | No | 0 | 4 | 3 |
| 425 | R:R:R153 | R:R:T152 | 1.29 | No | No | 0 | 5 | 5 |
| 426 | R:R:S76 | R:R:Y75 | 1.27 | No | No | 0 | 4 | 4 |
| 427 | R:R:V52 | R:R:Y49 | 1.26 | No | No | 0 | 5 | 2 |
| 428 | R:R:F50 | R:R:I53 | 1.26 | No | No | 0 | 5 | 5 |
| 429 | R:R:F117 | R:R:L118 | 1.22 | No | No | 0 | 1 | 6 |
| 430 | R:R:K48 | R:R:Y49 | 1.19 | No | No | 0 | 4 | 2 |
| Color | ConSurf Grade |
| No Conservation data available | |
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 | |
| 6 | |
| 7 | |
| 8 | |
| 9 |
Index: hub id, click on each number to highlight the corresponding hub in the 3D visualization.
Hub: the hub being considered.
Avg Int. Strength: the average interaction strength of all the links of the corresponding hub.
Num Of Links: the number of links of the corresponding hub.
Community: the id of the community the link belong to, otherwise 0.
ConSurf: this column reports the ConSurf conservation grades of each hub.
| Index | Hub | Avg Int. Strength | Num Of Links | Community | ConSurf |
|---|---|---|---|---|---|
| 1 | L:L:E40 | 5.81833 | 6 | 2 | 6 |
| 2 | L:L:R42 | 9.334 | 5 | 2 | 8 |
| 3 | L:L:Q44 | 5.2975 | 4 | 0 | 5 |
| 4 | L:L:K63 | 4.1425 | 4 | 1 | 5 |
| 5 | L:L:P67 | 4.46143 | 7 | 1 | 6 |
| 6 | L:L:H68 | 4.3475 | 4 | 1 | 8 |
| 7 | L:L:E73 | 6.655 | 4 | 7 | 9 |
| 8 | L:L:L78 | 5.7425 | 4 | 8 | 7 |
| 9 | L:L:L86 | 5.21 | 4 | 0 | 9 |
| 10 | L:L:N87 | 4.65 | 4 | 0 | 7 |
| 11 | L:L:S90 | 4.38 | 4 | 0 | 8 |
| 12 | L:L:K94 | 6.525 | 4 | 10 | 6 |
| 13 | L:L:L101 | 5 | 4 | 0 | 6 |
| 14 | N:N:I52 | 4.3375 | 4 | 0 | 8 |
| 15 | N:N:E73 | 5.272 | 5 | 11 | 9 |
| 16 | N:N:L86 | 4.8375 | 4 | 0 | 9 |
| 17 | N:N:N87 | 5.3825 | 4 | 0 | 7 |
| 18 | N:N:S90 | 4.7825 | 4 | 0 | 8 |
| 19 | R:R:Y55 | 5.79 | 4 | 0 | 7 |
| 20 | R:R:Y86 | 5.184 | 5 | 0 | 7 |
| 21 | R:R:L90 | 5.1475 | 4 | 12 | 9 |
| 22 | R:R:D94 | 7.515 | 4 | 12 | 9 |
| 23 | R:R:T100 | 3.5925 | 4 | 13 | 7 |
| 24 | R:R:W112 | 5.96 | 6 | 5 | 9 |
| 25 | R:R:F114 | 3.485 | 4 | 0 | 6 |
| 26 | R:R:N129 | 4.8025 | 4 | 0 | 8 |
| 27 | R:R:F130 | 6.7725 | 4 | 3 | 7 |
| 28 | R:R:Y131 | 8.83 | 4 | 0 | 7 |
| 29 | R:R:I134 | 4.8275 | 4 | 3 | 8 |
| 30 | R:R:Y145 | 5.93667 | 6 | 6 | 8 |
| 31 | R:R:V149 | 5.6575 | 4 | 6 | 7 |
| 32 | R:R:H150 | 6.53 | 4 | 6 | 6 |
| 33 | R:R:R184 | 9.91 | 4 | 0 | 5 |
| 34 | R:R:T186 | 4.1725 | 4 | 5 | 4 |
| 35 | R:R:R212 | 7.9025 | 4 | 2 | 5 |
| 36 | R:R:F220 | 6.825 | 4 | 3 | 8 |
| 37 | R:R:L224 | 4.205 | 4 | 3 | 7 |
| 38 | R:R:Y231 | 8.352 | 5 | 0 | 9 |
| 39 | R:R:T234 | 3.63 | 4 | 6 | 8 |
| 40 | R:R:M250 | 4.41 | 4 | 0 | 5 |
| 41 | R:R:V256 | 3.6925 | 4 | 4 | 8 |
| 42 | R:R:F260 | 5.428 | 5 | 3 | 9 |
| 43 | R:R:W264 | 5.30167 | 6 | 3 | 8 |
| 44 | R:R:Y267 | 4.57 | 4 | 0 | 7 |
| 45 | R:R:L271 | 6.8575 | 4 | 0 | 6 |
| 46 | R:R:D274 | 8.475 | 4 | 2 | 4 |
| 47 | R:R:M277 | 3.74 | 4 | 0 | 4 |
| 48 | R:R:R278 | 7.838 | 5 | 2 | 4 |
| 49 | R:R:E284 | 6.67 | 4 | 2 | 3 |
| 50 | R:R:R289 | 7.815 | 4 | 2 | 3 |
| 51 | R:R:Y314 | 6.306 | 5 | 4 | 9 |
| 52 | R:R:F316 | 4.415 | 4 | 0 | 6 |
| Color | ConSurf Grade |
| No Conservation data available | |
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 | |
| 6 | |
| 7 | |
| 8 | |
| 9 |
Index: link id, click on each number to highlight the corresponding link in the 3D visualization.
Node1 Node2: the two nodes of the corresponding link.
Recurrence: the relative Recurrence in the pool of shortest paths.
Int. Strength: the interaction strength between the two nodes.
Hub1?, Hub2?: "Yes" if the corresponding node has more than 3 links, otherwise "No".
Community: the id of the community the link belong to, otherwise 0.
ConSurf1, ConSurf2: these columns report the ConSurf conservation grades of the two nodes involved in a link.
| Index | Node1 | Node2 | Recurrence | Int. Strength | Hub1? | Hub2? | Community | ConSurf1 | ConSurf2 |
|---|---|---|---|---|---|---|---|---|---|
| 1 | L:L:E40 | R:R:R208 | 28.1191 | 10.47 | Yes | No | 0 | 6 | 4 |
| 2 | R:R:E198 | R:R:R208 | 29.062 | 10.47 | No | No | 0 | 5 | 4 |
| 3 | R:R:E198 | R:R:R184 | 29.2811 | 19.77 | No | Yes | 0 | 5 | 5 |
| 4 | R:R:C196 | R:R:R184 | 100 | 2.79 | No | Yes | 0 | 9 | 5 |
| 5 | R:R:C196 | R:R:W112 | 51.8484 | 7.84 | No | Yes | 5 | 9 | 9 |
| 6 | R:R:T186 | R:R:W112 | 48.4264 | 4.85 | Yes | Yes | 5 | 4 | 9 |
| 7 | R:R:T186 | R:R:Y188 | 93.2755 | 3.75 | Yes | No | 0 | 4 | 1 |
| 8 | R:R:C119 | R:R:C196 | 47.8969 | 7.28 | No | No | 5 | 9 | 9 |
| 9 | R:R:C119 | R:R:T186 | 47.5167 | 3.38 | No | Yes | 5 | 9 | 4 |
| 10 | L:L:E40 | L:L:R42 | 10.3514 | 4.65 | Yes | Yes | 2 | 6 | 8 |
| 11 | L:L:E40 | R:R:R212 | 39.5876 | 3.49 | Yes | Yes | 2 | 6 | 5 |
| 12 | L:L:E40 | R:R:R278 | 12.9455 | 6.98 | Yes | Yes | 2 | 6 | 4 |
| 13 | R:R:R208 | R:R:Y197 | 25.3912 | 18.52 | No | No | 0 | 4 | 3 |
| 14 | R:R:R278 | R:R:Y197 | 26.3791 | 3.09 | Yes | No | 0 | 4 | 3 |
| 15 | L:L:H68 | L:L:L41 | 48.7936 | 3.86 | Yes | No | 0 | 8 | 6 |
| 16 | L:L:L41 | R:R:Y197 | 49.7755 | 19.93 | No | No | 0 | 6 | 3 |
| 17 | L:L:R42 | R:R:R289 | 15.2222 | 10.66 | Yes | Yes | 2 | 8 | 3 |
| 18 | L:L:K63 | L:L:Q44 | 20.3594 | 4.07 | Yes | Yes | 0 | 5 | 5 |
| 19 | L:L:H68 | L:L:K63 | 28.6687 | 5.24 | Yes | Yes | 1 | 8 | 5 |
| 20 | R:R:E284 | R:R:R289 | 11.0396 | 5.82 | Yes | Yes | 2 | 3 | 3 |
| 21 | L:L:N87 | L:L:S90 | 11.0479 | 4.47 | Yes | Yes | 0 | 7 | 8 |
| 22 | L:L:S90 | L:L:V93 | 21.7322 | 4.85 | Yes | No | 0 | 8 | 8 |
| 23 | L:L:I97 | L:L:V93 | 36.3254 | 3.07 | No | No | 0 | 7 | 8 |
| 24 | L:L:I97 | L:L:L101 | 38.377 | 2.85 | No | Yes | 0 | 7 | 6 |
| 25 | L:L:L101 | N:N:V62 | 47.705 | 2.98 | Yes | No | 0 | 6 | 7 |
| 26 | N:N:V62 | N:N:V74 | 48.7107 | 6.41 | No | No | 0 | 7 | 8 |
| 27 | N:N:L86 | N:N:V74 | 49.7139 | 2.98 | Yes | No | 0 | 9 | 8 |
| 28 | N:N:I52 | N:N:L86 | 78.6184 | 4.28 | Yes | Yes | 0 | 8 | 9 |
| 29 | N:N:I52 | N:N:L78 | 84.2543 | 4.28 | Yes | No | 0 | 8 | 7 |
| 30 | N:N:L78 | N:N:N80 | 85.1699 | 8.24 | No | No | 0 | 7 | 5 |
| 31 | N:N:G81 | N:N:N80 | 86.9941 | 1.7 | No | No | 0 | 7 | 5 |
| 32 | N:N:G81 | R:R:Y188 | 87.8742 | 5.79 | No | No | 0 | 7 | 1 |
| 33 | L:L:L86 | L:L:V93 | 14.2734 | 8.94 | Yes | No | 0 | 9 | 8 |
| 34 | L:L:H68 | L:L:P67 | 18.2225 | 4.58 | Yes | Yes | 1 | 8 | 6 |
| 35 | N:N:L86 | N:N:V93 | 30.2465 | 8.94 | Yes | No | 0 | 9 | 8 |
| 36 | N:N:S90 | N:N:V93 | 29.1982 | 6.46 | Yes | No | 0 | 8 | 8 |
| 37 | N:N:N87 | N:N:S90 | 20.7266 | 4.47 | Yes | Yes | 0 | 7 | 8 |
| 38 | N:N:N87 | N:N:T48 | 11.0148 | 5.85 | Yes | No | 0 | 7 | 8 |
| 39 | R:R:R184 | R:R:V180 | 73.9514 | 6.54 | Yes | No | 0 | 5 | 4 |
| 40 | R:R:E127 | R:R:V180 | 73.9668 | 12.84 | No | No | 0 | 4 | 4 |
| 41 | R:R:E127 | R:R:Y131 | 74.0616 | 8.98 | No | Yes | 0 | 4 | 7 |
| 42 | R:R:F130 | R:R:Y131 | 74.4525 | 9.28 | Yes | Yes | 0 | 7 | 7 |
| 43 | R:R:F130 | R:R:W264 | 49.438 | 10.02 | Yes | Yes | 3 | 7 | 8 |
| 44 | R:R:W264 | R:R:Y267 | 49.4735 | 2.89 | Yes | Yes | 0 | 8 | 7 |
| 45 | R:R:E300 | R:R:Y267 | 18.1159 | 3.37 | No | Yes | 0 | 5 | 7 |
| 46 | R:R:E300 | R:R:I304 | 15.3371 | 4.1 | No | No | 0 | 5 | 7 |
| 47 | R:R:I304 | R:R:Y55 | 12.1258 | 8.46 | No | Yes | 0 | 7 | 7 |
| 48 | R:R:L271 | R:R:R212 | 57.1254 | 10.93 | Yes | Yes | 0 | 6 | 5 |
| 49 | R:R:L271 | R:R:Y267 | 51.6482 | 7.03 | Yes | Yes | 0 | 6 | 7 |
| 50 | R:R:H306 | R:R:W264 | 38.4765 | 7.41 | No | Yes | 0 | 9 | 8 |
| 51 | R:R:H306 | R:R:S307 | 27.9343 | 2.79 | No | No | 0 | 9 | 8 |
| 52 | R:R:D94 | R:R:S307 | 28.298 | 4.42 | Yes | No | 0 | 9 | 8 |
| 53 | R:R:D94 | R:R:N66 | 25.1578 | 6.73 | Yes | No | 0 | 9 | 9 |
| 54 | R:R:A91 | R:R:N66 | 20.7799 | 3.13 | No | No | 0 | 8 | 9 |
| 55 | R:R:A91 | R:R:V69 | 20.0443 | 3.39 | No | No | 0 | 8 | 8 |
| 56 | R:R:F316 | R:R:V69 | 19.3087 | 3.93 | Yes | No | 0 | 6 | 8 |
| 57 | R:R:A315 | R:R:F316 | 21.7938 | 2.77 | No | Yes | 0 | 7 | 6 |
| 58 | R:R:A315 | R:R:F321 | 19.0221 | 8.32 | No | No | 0 | 7 | 8 |
| 59 | R:R:F321 | R:R:V72 | 10.5789 | 5.24 | No | No | 0 | 8 | 7 |
| 60 | R:R:F130 | R:R:I134 | 25.5606 | 6.28 | Yes | Yes | 3 | 7 | 8 |
| 61 | R:R:F260 | R:R:I134 | 24.8913 | 3.77 | Yes | Yes | 3 | 9 | 8 |
| 62 | R:R:F260 | R:R:M227 | 70.8279 | 11.2 | Yes | No | 0 | 9 | 8 |
| 63 | R:R:L137 | R:R:M227 | 33.3179 | 5.65 | No | No | 0 | 8 | 8 |
| 64 | R:R:L137 | R:R:Y314 | 30.6445 | 9.38 | No | Yes | 0 | 8 | 9 |
| 65 | R:R:I140 | R:R:Y314 | 16.1271 | 6.04 | No | Yes | 4 | 9 | 9 |
| 66 | R:R:I140 | R:R:Y86 | 16.2527 | 6.04 | No | Yes | 0 | 9 | 7 |
| 67 | R:R:F260 | R:R:W264 | 50.8771 | 3.01 | Yes | Yes | 3 | 9 | 8 |
| 68 | R:R:L137 | R:R:L90 | 14.9663 | 5.54 | No | Yes | 0 | 8 | 9 |
| 69 | R:R:D94 | R:R:L90 | 10.3171 | 6.79 | Yes | Yes | 12 | 9 | 9 |
| 70 | R:R:F97 | R:R:H306 | 14.3575 | 9.05 | No | No | 0 | 6 | 9 |
| 71 | R:R:M227 | R:R:S141 | 43.7914 | 4.6 | No | No | 0 | 8 | 9 |
| 72 | R:R:S141 | R:R:Y231 | 25.7679 | 13.99 | No | Yes | 0 | 9 | 9 |
| 73 | R:R:C230 | R:R:S141 | 18.2154 | 3.44 | No | No | 0 | 7 | 9 |
| 74 | R:R:C230 | R:R:Y145 | 16.3368 | 6.72 | No | Yes | 0 | 7 | 8 |
| 75 | R:R:I253 | R:R:Y231 | 21.2253 | 9.67 | No | Yes | 0 | 7 | 9 |
| 76 | R:R:I253 | R:R:L235 | 15.1298 | 7.14 | No | No | 0 | 7 | 4 |
| 77 | R:R:L235 | R:R:M250 | 12.82 | 2.83 | No | Yes | 0 | 4 | 5 |
| 78 | L:L:P67 | R:R:T204 | 12.9621 | 1.75 | Yes | No | 0 | 6 | 3 |
| 79 | R:R:M200 | R:R:T204 | 10.8252 | 3.01 | No | No | 0 | 5 | 3 |
| 80 | R:R:V252 | R:R:V256 | 11.2078 | 1.6 | No | Yes | 0 | 7 | 8 |
| 81 | R:R:D274 | R:R:R212 | 17.7392 | 10.72 | Yes | Yes | 2 | 4 | 5 |
| 82 | R:R:D274 | R:R:R278 | 12.377 | 8.34 | Yes | Yes | 2 | 4 | 4 |
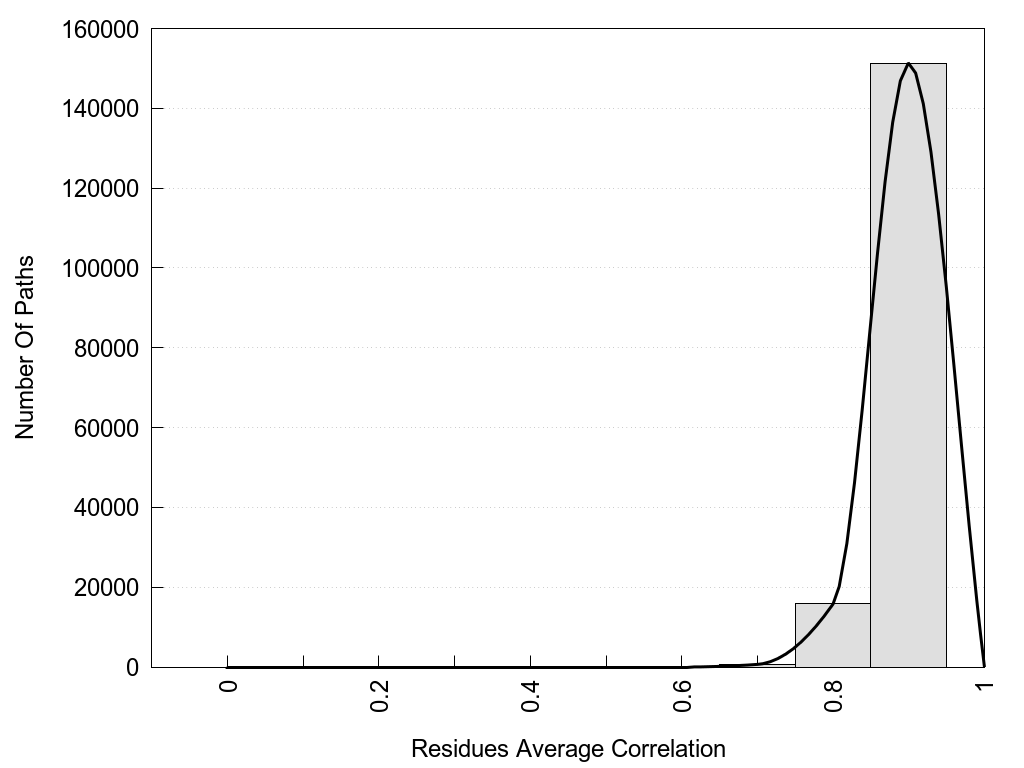
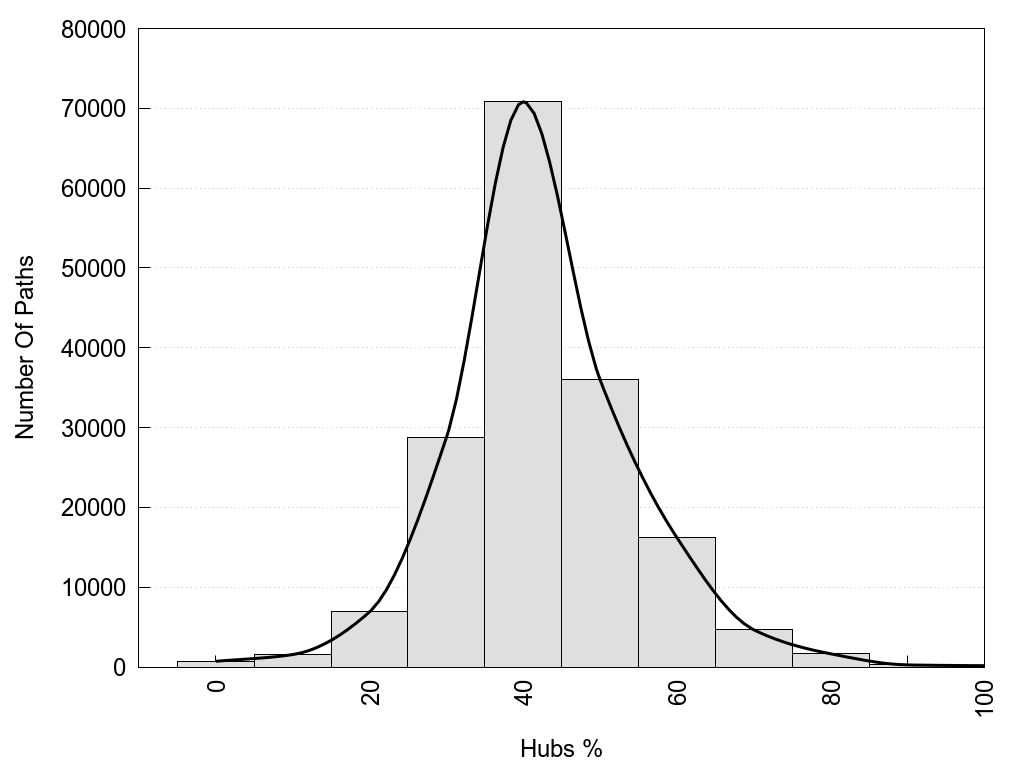
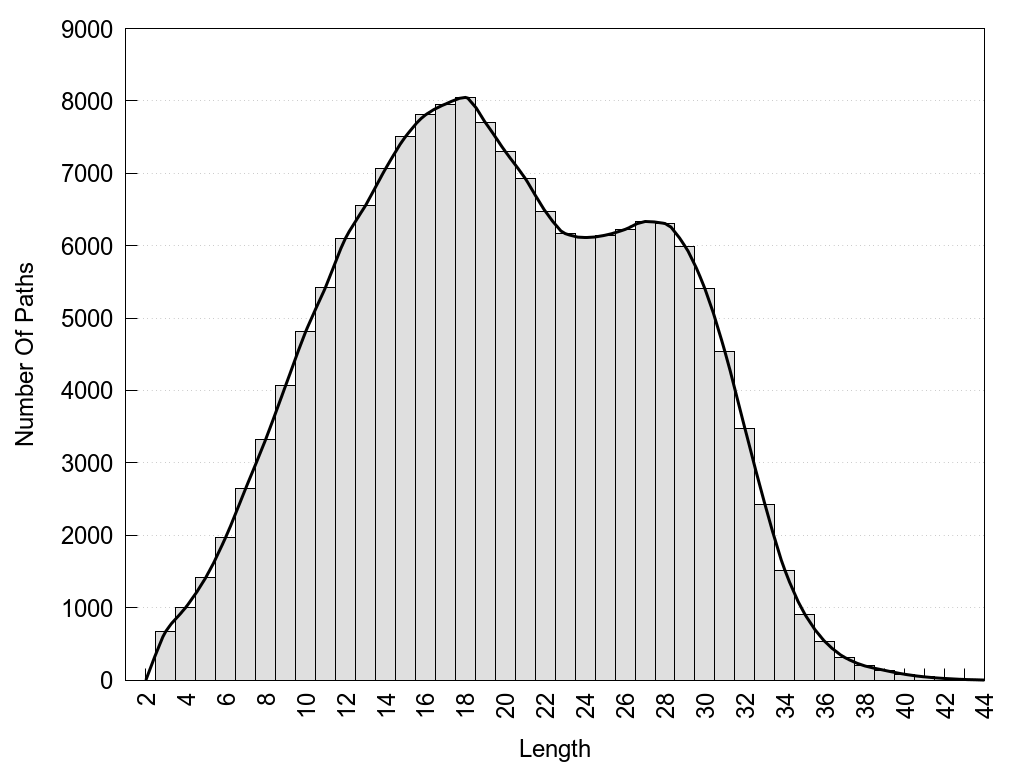
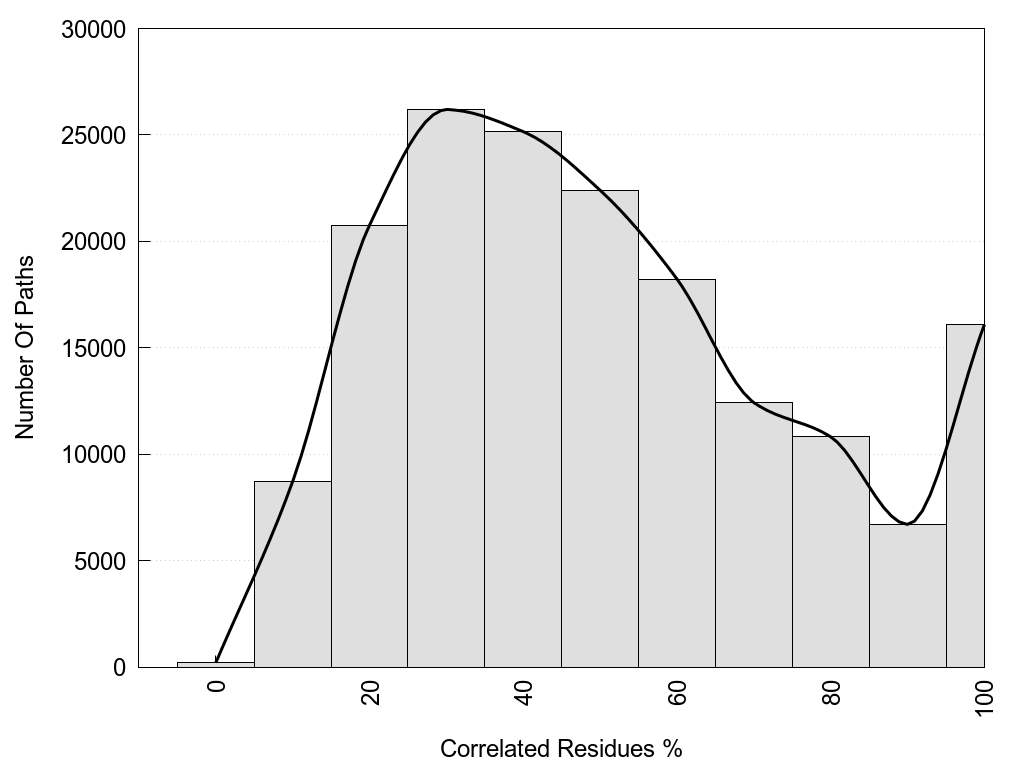
2D representation of the global metapath, ligand(s) interactions and
histograms of path distribution according to several parameters
(click on the image to enlarge it 🔍):
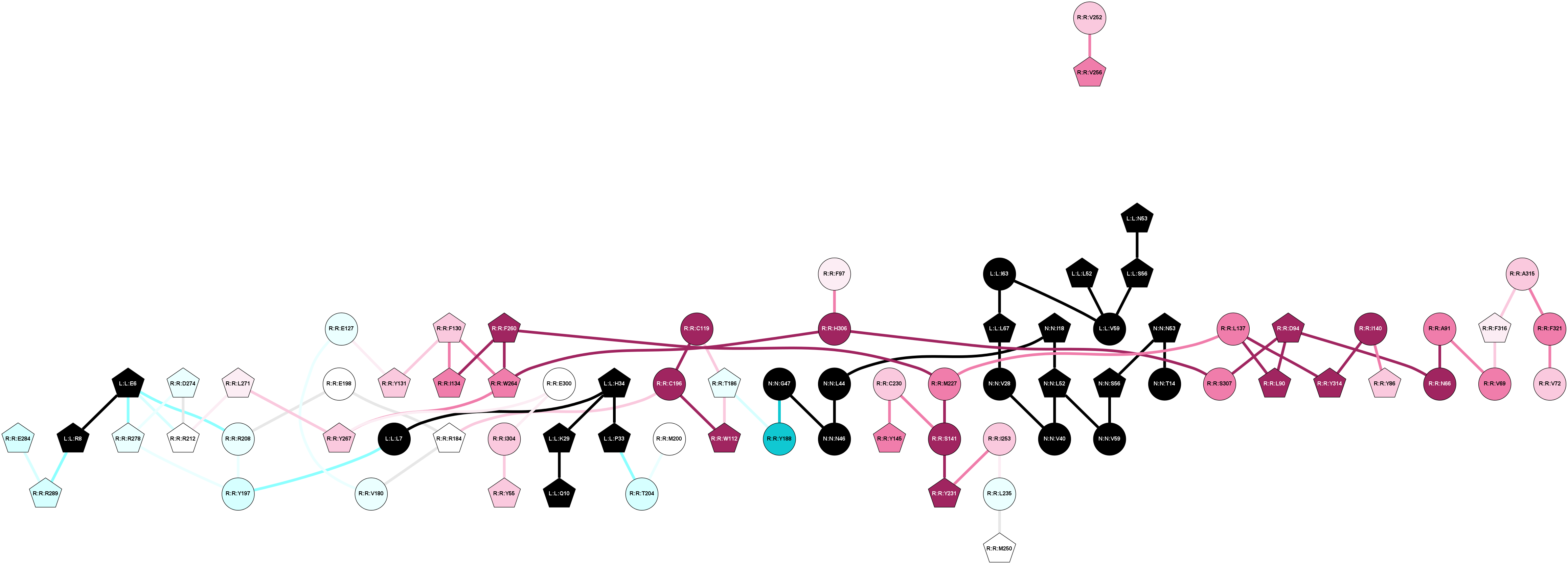
A 2D representation of the global communication in the network.
ConSurf Conservation Grade (See documentation):
n/a 1 2 3 4 5 6 7 8 9
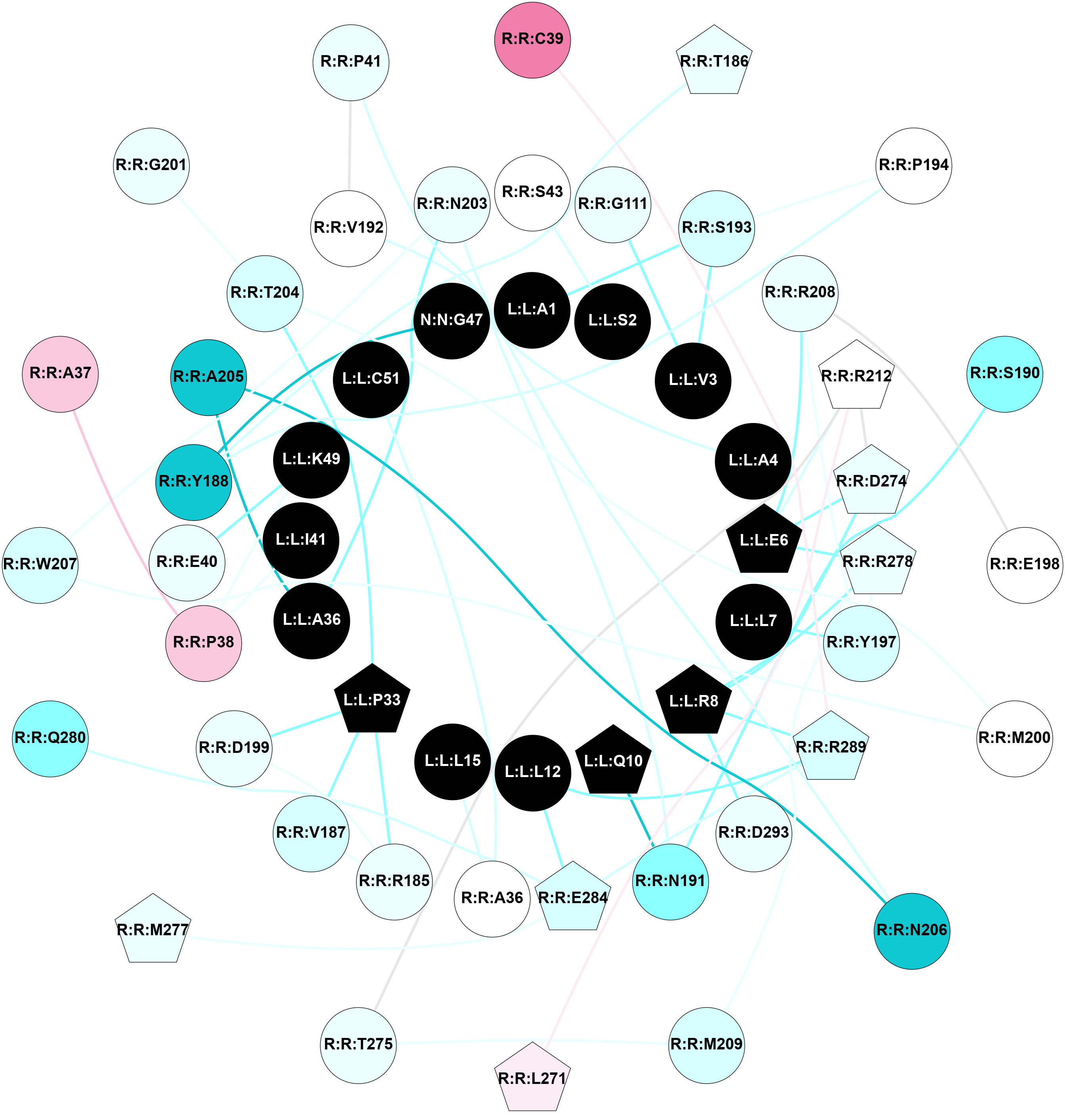
2D representation of the interactions of this orthosteric/allosteric ligand. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Links and nodes colored according to ConSurf Conservation Grade (See documentation): n/a 1 2 3 4 5 6 7 8 9 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Location and physicochemical properties of the interaction partners of this ligand | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Interactions of this ligand | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Similarities between the interactions of this ligand and those of other networks | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| Annotation | Type | Links |
|---|---|---|
| Gene Ontology | Molecular Function | |
| Gene Ontology | Biological Process | |
| Gene Ontology | Cellular Component | |
| SCOP2 | Domain Identifier | • Transducin (heterotrimeric G protein), gamma chain • G protein-coupled receptor-like • Interleukin 8-like chemokines |
| SCOP2 | Family Identifier | • Transducin (heterotrimeric G protein), gamma chain • G protein-coupled receptor-like • Interleukin 8-like chemokines |
| Membrane Protein Annotations | - | • Orientations of Proteins in Membranes database (OPM) • Protein Data Bank of Transmembrane Proteins (PDBTM) • MemProtMD |
Details about the values in these tables can be found in the corresponding documentation page .
| PDBsum | Open PDBsum Page |
| Chain | L |
| Protein | CXCL1 |
| UniProt | P09341 |
| Sequence | >8XWV_nogp_Chain_L ASVATELRC QCLQTLQGI HPKNIQSVN VKSPGPHCA QTEVIATLK NGRKACLNP ASPIVKKII EKMLN Click on each residue to open a popup with some information about it. ConSurf Conservation Grade (See documentation): n/a 1 2 3 4 5 6 7 8 9 |
| Chain | N |
| Protein | CXCL1 |
| UniProt | P09341 |
| Sequence | >8XWV_nogp_Chain_N CQCLQTLQG IHPKNIQSV NVKSPGPHC AQTEVIATL KNGRKACLN PASPIVKKI IEKMLNS Click on each residue to open a popup with some information about it. ConSurf Conservation Grade (See documentation): n/a 1 2 3 4 5 6 7 8 9 |
| Chain | R |
| Protein | Receptor |
| UniProt | P25025 |
| Sequence | >8XWV_nogp_Chain_R DAAPCEPES LEINKYFVV IIYALVFLL SLLGNSLVM LVILYSRVG RSVTDVYLL NLALADLLF ALTLPIWAA SKVNGWIFG TFLCKVVSL LKEVNFYSG ILLLACISV DRYLAIVHA TRTLTQKRY LVKFICLSI WGLSLLLAL PVLLFRRTV YSSNVSPAC YEDMGNNTA NWRMLLRIL PQSFGFIVP LLIMLFCYG FTLRTLFKA HMGQKHRAM RVIFAVVLI FLLCWLPYN LVLLADTLM RTQVIQETC ERRNHIDRA LDATEILGI LHSCLNPLI YAFIGQKFR HGLLKILA Click on each residue to open a popup with some information about it. ConSurf Conservation Grade (See documentation): n/a 1 2 3 4 5 6 7 8 9 |
| This receptor, from the same or other species and bound to the same or other ligands, is also present in the following networks: | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Show | PDB | Class | SubFamily | Type | SubType | Species | Orthosteric Ligand | Other Ligand(s) | Protein Partners | Resolution | Date | DOI |
| 6LFL | A | Protein | Chemokine | CXCR2 | Homo sapiens | - | PubChem 153466996 | - | 3.2 | 2020-09-02 | doi.org/10.1038/s41586-020-2492-5 | |
| 6LFM | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL8 | - | Gi1/β1/γ2 | 3.5 | 2020-09-02 | doi.org/10.1038/s41586-020-2492-5 | |
| 6LFM (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL8 | - | 3.5 | 2020-09-02 | doi.org/10.1038/s41586-020-2492-5 | ||
| 6LFO | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL8 | - | Gi1/β1/γ2 | 3.4 | 2020-09-02 | doi.org/10.1038/s41586-020-2492-5 | |
| 6LFO (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL8 | - | 3.4 | 2020-09-02 | doi.org/10.1038/s41586-020-2492-5 | ||
| 8XVU | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL2 | - | - | 3.09 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XWA | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL1 | - | - | 3.48 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XWF | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL3 | - | - | 3.65 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XWM | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL6 | - | - | 3.71 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XWN | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL8 | - | - | 3.29 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XWS | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL5 | - | - | 3.06 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XWV | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL1 | - | Go/β1/γ2 | 3.07 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XWV (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL1 | - | 3.07 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | ||
| 8XX3 | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL3 | - | Go/β1/γ2 | 3.38 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XX3 (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL3 | - | 3.38 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | ||
| 8XX6 | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL8 | - | Go/β1/γ2 | 2.99 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XX6 (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL8 | - | 2.99 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | ||
| 8XX7 | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL5 | - | Go/β1/γ2 | 3.32 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XX7 (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL5 | - | 3.32 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | ||
| 8XXH | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL2 | - | Go/β1/γ2 | 2.8 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XXH (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL2 | - | 2.8 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | ||
| 8XXR | A | Protein | Chemokine | CXCR2 | Homo sapiens | - | - | Go/β1/γ2 | 3.17 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XXR (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | - | - | 3.17 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | ||
| 8XXX | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL6 | - | Go/β1/γ2 | 3.17 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | |
| 8XXX (No Gprot) | A | Protein | Chemokine | CXCR2 | Homo sapiens | CXCL6 | - | 3.17 | 2025-01-15 | doi.org/10.1016/j.molcel.2025.01.024 | ||
You can download a compressed (zip) file with structure(s), 3D outputs (as PyMol and VMD scripts) and numerical data files (as csv and plain text files).
Download 8XWV_nogp.zipYou can click to copy the link of this page to easily come back here later
or use this QR code to link and share this page.
You can also read or download a guide explaining the meaning of all files and numerical data.