- Press and hold down the left mouse button and move the mouse to rotate.
- Press and hold down the central mouse button and move the mouse to translate.
- Use the mouse wheel to zoom in or out.
- Click on any atom on your molecule and the corresponding residue label will be placed beneath the viewer.
| Color | ConSurf Grade |
| No Conservation data available | |
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 | |
| 6 | |
| 7 | |
| 8 | |
| 9 |
Index: link id, click on each number to highlight the corresponding link in the 3D visualization.
Node1 Node2: the two nodes of the corresponding link.
Int. Strength: the interaction strength between the two nodes.
Hub1?, Hub2?: "Yes" if the corresponding node has more than 3 links, otherwise "No".
Community: the id of the community the link belong to, otherwise 0.
ConSurf1, ConSurf2: these columns report the ConSurf conservation grades of the two nodes involved in a link.
| Index | Node1 | Node2 | Int. Strength | Hub1? | Hub2? | Community | ConSurf1 | ConSurf2 |
|---|---|---|---|---|---|---|---|---|
| 1 | L:L:?1 | R:R:D116 | 18.59 | Yes | Yes | 1 | 0 | 8 |
| 2 | L:L:?1 | R:R:V117 | 8.59 | Yes | No | 0 | 0 | 8 |
| 3 | L:L:?1 | R:R:C120 | 4.58 | Yes | No | 1 | 0 | 7 |
| 4 | L:L:?1 | R:R:T121 | 4.25 | Yes | Yes | 0 | 0 | 8 |
| 5 | L:L:?1 | R:R:I189 | 7.2 | Yes | No | 0 | 0 | 6 |
| 6 | L:L:?1 | R:R:S199 | 6.49 | Yes | No | 0 | 0 | 8 |
| 7 | L:L:?1 | R:R:F361 | 25.46 | Yes | Yes | 1 | 0 | 8 |
| 8 | L:L:?1 | R:R:F362 | 8.78 | Yes | Yes | 1 | 0 | 7 |
| 9 | L:L:?1 | R:R:Y390 | 14.37 | Yes | Yes | 1 | 0 | 8 |
| 10 | R:R:Q36 | R:R:V98 | 5.73 | No | Yes | 0 | 4 | 6 |
| 11 | R:R:S40 | R:R:V98 | 6.46 | No | Yes | 0 | 5 | 6 |
| 12 | R:R:L43 | R:R:W387 | 18.22 | No | No | 0 | 7 | 8 |
| 13 | R:R:G44 | R:R:I47 | 3.53 | No | No | 0 | 4 | 7 |
| 14 | R:R:I47 | R:R:L90 | 7.14 | No | No | 0 | 7 | 8 |
| 15 | R:R:L83 | R:R:V51 | 4.47 | Yes | No | 0 | 8 | 7 |
| 16 | R:R:G53 | R:R:Y402 | 4.35 | No | Yes | 0 | 9 | 6 |
| 17 | R:R:D82 | R:R:N54 | 8.08 | Yes | Yes | 0 | 9 | 9 |
| 18 | R:R:L83 | R:R:N54 | 4.12 | Yes | Yes | 0 | 8 | 9 |
| 19 | R:R:N54 | R:R:P397 | 6.52 | Yes | No | 0 | 9 | 9 |
| 20 | R:R:C56 | R:R:Y402 | 4.03 | No | Yes | 0 | 8 | 6 |
| 21 | R:R:P397 | R:R:V57 | 3.53 | No | Yes | 0 | 9 | 9 |
| 22 | R:R:A401 | R:R:V57 | 5.09 | No | Yes | 0 | 8 | 9 |
| 23 | R:R:V57 | R:R:Y402 | 5.05 | Yes | Yes | 0 | 9 | 6 |
| 24 | R:R:V58 | R:R:V80 | 6.41 | No | No | 0 | 7 | 7 |
| 25 | R:R:A60 | R:R:F411 | 5.55 | No | No | 0 | 8 | 8 |
| 26 | R:R:I61 | R:R:L67 | 4.28 | No | Yes | 4 | 8 | 8 |
| 27 | R:R:F407 | R:R:I61 | 6.28 | Yes | No | 4 | 8 | 8 |
| 28 | R:R:E64 | R:R:K413 | 9.45 | No | No | 0 | 7 | 6 |
| 29 | R:R:E64 | R:R:I414 | 8.2 | No | No | 0 | 7 | 7 |
| 30 | R:R:L67 | R:R:N72 | 6.87 | Yes | No | 4 | 8 | 9 |
| 31 | R:R:F407 | R:R:L67 | 6.09 | Yes | Yes | 4 | 8 | 8 |
| 32 | R:R:Q68 | R:R:Y73 | 6.76 | No | No | 0 | 8 | 6 |
| 33 | R:R:N69 | R:R:N72 | 10.9 | No | No | 0 | 8 | 9 |
| 34 | R:R:R148 | R:R:V70 | 14.38 | Yes | No | 0 | 8 | 7 |
| 35 | R:R:P150 | R:R:V70 | 5.3 | No | No | 0 | 5 | 7 |
| 36 | R:R:F407 | R:R:N72 | 12.08 | Yes | No | 4 | 8 | 9 |
| 37 | R:R:P150 | R:R:Y73 | 5.56 | No | No | 0 | 5 | 6 |
| 38 | R:R:D133 | R:R:L74 | 5.43 | No | No | 0 | 9 | 8 |
| 39 | R:R:L156 | R:R:L74 | 5.54 | No | No | 0 | 6 | 8 |
| 40 | R:R:I75 | R:R:Y400 | 6.04 | No | Yes | 0 | 8 | 9 |
| 41 | R:R:H126 | R:R:S77 | 6.97 | Yes | No | 1 | 8 | 9 |
| 42 | R:R:I157 | R:R:S77 | 10.84 | No | No | 1 | 8 | 9 |
| 43 | R:R:D82 | R:R:L78 | 8.14 | Yes | Yes | 1 | 9 | 9 |
| 44 | R:R:H126 | R:R:L78 | 6.43 | Yes | Yes | 1 | 8 | 9 |
| 45 | R:R:L127 | R:R:L78 | 5.54 | Yes | Yes | 1 | 8 | 9 |
| 46 | R:R:I130 | R:R:L78 | 4.28 | No | Yes | 1 | 9 | 9 |
| 47 | R:R:L78 | R:R:N396 | 12.36 | Yes | Yes | 1 | 9 | 9 |
| 48 | R:R:L78 | R:R:Y400 | 4.69 | Yes | Yes | 1 | 9 | 9 |
| 49 | R:R:S122 | R:R:T81 | 6.4 | No | No | 1 | 8 | 9 |
| 50 | R:R:H126 | R:R:T81 | 12.32 | Yes | No | 1 | 8 | 9 |
| 51 | R:R:T81 | R:R:W161 | 9.7 | No | Yes | 1 | 9 | 9 |
| 52 | R:R:D82 | R:R:S123 | 5.89 | Yes | No | 0 | 9 | 9 |
| 53 | R:R:D82 | R:R:S393 | 4.42 | Yes | No | 0 | 9 | 9 |
| 54 | R:R:D82 | R:R:N396 | 9.42 | Yes | Yes | 1 | 9 | 9 |
| 55 | R:R:M84 | R:R:W161 | 4.65 | No | Yes | 0 | 7 | 9 |
| 56 | R:R:S393 | R:R:V85 | 6.46 | No | No | 0 | 9 | 8 |
| 57 | R:R:L115 | R:R:L88 | 5.54 | No | No | 0 | 5 | 6 |
| 58 | R:R:D116 | R:R:V89 | 7.3 | Yes | No | 1 | 8 | 8 |
| 59 | R:R:V89 | R:R:Y390 | 3.79 | No | Yes | 1 | 8 | 8 |
| 60 | R:R:L90 | R:R:Y390 | 8.21 | No | Yes | 0 | 8 | 8 |
| 61 | R:R:M92 | R:R:W102 | 4.65 | No | Yes | 2 | 6 | 9 |
| 62 | R:R:F112 | R:R:M92 | 12.44 | No | No | 2 | 6 | 6 |
| 63 | R:R:A93 | R:R:Y390 | 4 | No | Yes | 0 | 7 | 8 |
| 64 | R:R:L95 | R:R:L99 | 5.54 | No | No | 0 | 6 | 5 |
| 65 | R:R:W102 | R:R:Y96 | 11.58 | Yes | Yes | 2 | 9 | 6 |
| 66 | R:R:C187 | R:R:Y96 | 4.03 | Yes | Yes | 2 | 9 | 6 |
| 67 | R:R:A383 | R:R:Q97 | 4.55 | No | No | 0 | 6 | 7 |
| 68 | R:R:Q97 | R:R:W387 | 8.76 | No | No | 0 | 7 | 8 |
| 69 | R:R:L99 | R:R:T103 | 7.37 | No | No | 0 | 5 | 5 |
| 70 | R:R:K101 | R:R:T103 | 6.01 | No | No | 0 | 5 | 5 |
| 71 | R:R:D185 | R:R:K101 | 16.59 | No | No | 0 | 4 | 5 |
| 72 | R:R:L104 | R:R:W102 | 10.25 | No | Yes | 2 | 7 | 9 |
| 73 | R:R:T108 | R:R:W102 | 4.85 | No | Yes | 2 | 6 | 9 |
| 74 | R:R:C109 | R:R:W102 | 6.53 | Yes | Yes | 2 | 9 | 9 |
| 75 | R:R:F112 | R:R:W102 | 9.02 | No | Yes | 2 | 6 | 9 |
| 76 | R:R:C187 | R:R:W102 | 10.45 | Yes | Yes | 2 | 9 | 9 |
| 77 | R:R:L104 | R:R:T108 | 8.84 | No | No | 2 | 7 | 6 |
| 78 | R:R:D110 | R:R:Q106 | 11.75 | No | No | 0 | 7 | 4 |
| 79 | R:R:C109 | R:R:C187 | 7.28 | Yes | Yes | 2 | 9 | 9 |
| 80 | R:R:L111 | R:R:L115 | 6.92 | No | No | 0 | 5 | 5 |
| 81 | R:R:C120 | R:R:D116 | 4.67 | No | Yes | 1 | 7 | 8 |
| 82 | R:R:D116 | R:R:Y390 | 11.49 | Yes | Yes | 1 | 8 | 8 |
| 83 | R:R:L118 | R:R:W161 | 6.83 | No | Yes | 0 | 6 | 9 |
| 84 | R:R:F165 | R:R:L118 | 14.61 | No | No | 0 | 5 | 6 |
| 85 | R:R:L118 | R:R:S168 | 4.5 | No | No | 0 | 6 | 8 |
| 86 | R:R:C120 | R:R:W358 | 5.22 | No | Yes | 1 | 7 | 8 |
| 87 | R:R:G164 | R:R:T121 | 3.64 | No | Yes | 0 | 8 | 8 |
| 88 | R:R:I167 | R:R:T121 | 6.08 | No | Yes | 0 | 8 | 8 |
| 89 | R:R:A203 | R:R:T121 | 5.03 | No | Yes | 0 | 8 | 8 |
| 90 | R:R:H126 | R:R:S122 | 6.97 | Yes | No | 1 | 8 | 8 |
| 91 | R:R:S122 | R:R:W161 | 19.77 | No | Yes | 1 | 8 | 9 |
| 92 | R:R:F354 | R:R:I124 | 5.02 | Yes | No | 1 | 9 | 8 |
| 93 | R:R:I124 | R:R:W358 | 12.92 | No | Yes | 1 | 8 | 8 |
| 94 | R:R:C128 | R:R:W125 | 3.92 | Yes | Yes | 1 | 7 | 7 |
| 95 | R:R:I163 | R:R:W125 | 19.97 | No | Yes | 0 | 5 | 7 |
| 96 | R:R:I206 | R:R:W125 | 3.52 | No | Yes | 0 | 6 | 7 |
| 97 | R:R:P207 | R:R:W125 | 16.21 | No | Yes | 1 | 9 | 7 |
| 98 | R:R:H126 | R:R:I157 | 7.95 | Yes | No | 1 | 8 | 8 |
| 99 | R:R:H126 | R:R:T160 | 20.54 | Yes | No | 1 | 8 | 8 |
| 100 | R:R:H126 | R:R:W161 | 5.29 | Yes | Yes | 1 | 8 | 9 |
| 101 | R:R:L127 | R:R:M211 | 5.65 | Yes | Yes | 1 | 8 | 9 |
| 102 | R:R:F354 | R:R:L127 | 4.87 | Yes | Yes | 1 | 9 | 8 |
| 103 | R:R:L127 | R:R:N396 | 12.36 | Yes | Yes | 1 | 8 | 9 |
| 104 | R:R:L127 | R:R:Y400 | 7.03 | Yes | Yes | 1 | 8 | 9 |
| 105 | R:R:C128 | R:R:P207 | 3.77 | Yes | No | 1 | 7 | 9 |
| 106 | R:R:C128 | R:R:M211 | 4.86 | Yes | Yes | 1 | 7 | 9 |
| 107 | R:R:A129 | R:R:L156 | 6.3 | No | No | 0 | 8 | 6 |
| 108 | R:R:I130 | R:R:R134 | 7.52 | No | Yes | 1 | 9 | 9 |
| 109 | R:R:I130 | R:R:Y400 | 10.88 | No | Yes | 1 | 9 | 9 |
| 110 | R:R:A131 | R:R:Y215 | 6.67 | No | Yes | 0 | 9 | 9 |
| 111 | R:R:L132 | R:R:W136 | 18.22 | No | No | 0 | 6 | 5 |
| 112 | R:R:L132 | R:R:L214 | 4.15 | No | No | 0 | 6 | 7 |
| 113 | R:R:D133 | R:R:R148 | 11.91 | No | Yes | 0 | 9 | 8 |
| 114 | R:R:R134 | R:R:Y215 | 8.23 | Yes | Yes | 1 | 9 | 9 |
| 115 | R:R:I350 | R:R:R134 | 3.76 | No | Yes | 1 | 8 | 9 |
| 116 | R:R:R134 | R:R:Y400 | 6.17 | Yes | Yes | 1 | 9 | 9 |
| 117 | R:R:W136 | R:R:Y135 | 4.82 | No | Yes | 0 | 5 | 9 |
| 118 | R:R:T139 | R:R:Y135 | 14.98 | No | Yes | 0 | 8 | 9 |
| 119 | R:R:L214 | R:R:Y135 | 5.86 | No | Yes | 0 | 7 | 9 |
| 120 | R:R:R217 | R:R:Y135 | 16.46 | No | Yes | 0 | 7 | 9 |
| 121 | R:R:I218 | R:R:Y135 | 3.63 | No | Yes | 0 | 9 | 9 |
| 122 | R:R:D140 | R:R:D143 | 9.31 | No | No | 0 | 7 | 6 |
| 123 | R:R:D143 | R:R:K147 | 4.15 | No | No | 0 | 6 | 5 |
| 124 | R:R:D143 | R:R:R152 | 11.91 | No | No | 0 | 6 | 7 |
| 125 | R:R:R148 | R:R:Y144 | 15.43 | Yes | No | 0 | 8 | 8 |
| 126 | R:R:R148 | R:R:R152 | 4.26 | Yes | No | 0 | 8 | 7 |
| 127 | R:R:T160 | R:R:W161 | 4.85 | No | Yes | 1 | 8 | 9 |
| 128 | R:R:L162 | R:R:L166 | 6.92 | No | No | 0 | 5 | 5 |
| 129 | R:R:I167 | R:R:Y198 | 4.84 | No | No | 0 | 8 | 5 |
| 130 | R:R:P170 | R:R:P171 | 5.84 | Yes | Yes | 3 | 8 | 7 |
| 131 | R:R:P170 | R:R:Y195 | 5.56 | Yes | Yes | 3 | 8 | 8 |
| 132 | R:R:P170 | R:R:Y198 | 5.56 | Yes | No | 0 | 8 | 5 |
| 133 | R:R:P171 | R:R:R176 | 4.32 | Yes | No | 3 | 7 | 6 |
| 134 | R:R:P171 | R:R:S190 | 3.56 | Yes | Yes | 3 | 7 | 6 |
| 135 | R:R:P171 | R:R:Y195 | 11.13 | Yes | Yes | 3 | 7 | 8 |
| 136 | R:R:D192 | R:R:W175 | 4.47 | No | No | 0 | 6 | 6 |
| 137 | R:R:R176 | R:R:S190 | 3.95 | No | Yes | 3 | 6 | 6 |
| 138 | R:R:I189 | R:R:Y195 | 7.25 | No | Yes | 0 | 6 | 8 |
| 139 | R:R:D192 | R:R:S190 | 7.36 | No | Yes | 0 | 6 | 6 |
| 140 | R:R:S190 | R:R:Y195 | 7.63 | Yes | Yes | 3 | 6 | 8 |
| 141 | R:R:F370 | R:R:H193 | 22.63 | No | No | 0 | 5 | 4 |
| 142 | R:R:E372 | R:R:H193 | 8.62 | No | No | 0 | 3 | 4 |
| 143 | R:R:S199 | R:R:Y195 | 11.45 | No | Yes | 0 | 8 | 8 |
| 144 | R:R:P369 | R:R:T196 | 12.24 | No | No | 0 | 7 | 7 |
| 145 | R:R:F370 | R:R:I197 | 3.77 | No | No | 0 | 5 | 7 |
| 146 | R:R:T200 | R:R:Y205 | 3.75 | Yes | Yes | 1 | 7 | 7 |
| 147 | R:R:F362 | R:R:T200 | 5.19 | Yes | Yes | 1 | 7 | 7 |
| 148 | R:R:L366 | R:R:T200 | 5.9 | No | Yes | 1 | 7 | 7 |
| 149 | R:R:F201 | R:R:Y205 | 8.25 | No | Yes | 1 | 5 | 7 |
| 150 | R:R:F201 | R:R:L366 | 4.87 | No | No | 1 | 5 | 7 |
| 151 | R:R:G202 | R:R:I206 | 3.53 | No | No | 0 | 5 | 6 |
| 152 | R:R:F204 | R:R:Y205 | 9.28 | Yes | Yes | 1 | 8 | 7 |
| 153 | R:R:F204 | R:R:L208 | 9.74 | Yes | No | 1 | 8 | 7 |
| 154 | R:R:F204 | R:R:F354 | 7.5 | Yes | Yes | 1 | 8 | 9 |
| 155 | R:R:F204 | R:R:W358 | 5.01 | Yes | Yes | 1 | 8 | 8 |
| 156 | R:R:F204 | R:R:L359 | 7.31 | Yes | No | 0 | 8 | 7 |
| 157 | R:R:F204 | R:R:F362 | 21.43 | Yes | Yes | 1 | 8 | 7 |
| 158 | R:R:L366 | R:R:Y205 | 5.86 | No | Yes | 1 | 7 | 7 |
| 159 | R:R:F354 | R:R:L208 | 3.65 | Yes | No | 1 | 9 | 7 |
| 160 | R:R:M211 | R:R:Y215 | 3.59 | Yes | Yes | 1 | 9 | 9 |
| 161 | R:R:M211 | R:R:M351 | 4.33 | Yes | Yes | 1 | 9 | 8 |
| 162 | R:R:F354 | R:R:M211 | 11.2 | Yes | Yes | 1 | 9 | 9 |
| 163 | R:R:L212 | R:R:M351 | 4.24 | No | Yes | 0 | 6 | 8 |
| 164 | R:R:R217 | R:R:V213 | 3.92 | No | No | 0 | 7 | 5 |
| 165 | R:R:I218 | R:R:Y215 | 7.25 | No | Yes | 0 | 9 | 9 |
| 166 | R:R:L347 | R:R:Y215 | 7.03 | No | Yes | 0 | 8 | 9 |
| 167 | R:R:I350 | R:R:Y215 | 8.46 | No | Yes | 1 | 8 | 9 |
| 168 | R:R:M351 | R:R:Y215 | 5.99 | Yes | Yes | 1 | 8 | 9 |
| 169 | R:R:R217 | R:R:R220 | 7.46 | No | No | 0 | 7 | 5 |
| 170 | R:R:F219 | R:R:R223 | 6.41 | Yes | No | 0 | 6 | 7 |
| 171 | R:R:F219 | R:R:L347 | 3.65 | Yes | No | 0 | 6 | 8 |
| 172 | R:R:A222 | R:R:T343 | 5.03 | No | No | 0 | 8 | 8 |
| 173 | R:R:R223 | R:R:R227 | 13.86 | No | No | 0 | 7 | 6 |
| 174 | R:R:F224 | R:R:K228 | 17.37 | No | No | 0 | 6 | 5 |
| 175 | R:R:K228 | R:R:R225 | 9.9 | No | No | 0 | 5 | 7 |
| 176 | R:R:I226 | R:R:R339 | 5.01 | No | No | 0 | 6 | 7 |
| 177 | R:R:E340 | R:R:I226 | 8.2 | No | No | 0 | 8 | 6 |
| 178 | R:R:E340 | R:R:R227 | 8.14 | No | No | 0 | 8 | 6 |
| 179 | R:R:R339 | R:R:T229 | 3.88 | No | No | 0 | 7 | 4 |
| 180 | R:R:E330 | R:R:R333 | 4.65 | No | No | 0 | 4 | 4 |
| 181 | R:R:M335 | R:R:R339 | 8.69 | No | No | 0 | 4 | 7 |
| 182 | R:R:I349 | R:R:K345 | 4.36 | No | No | 0 | 7 | 7 |
| 183 | R:R:F403 | R:R:I349 | 3.77 | No | No | 0 | 8 | 7 |
| 184 | R:R:L395 | R:R:T353 | 4.42 | No | No | 0 | 6 | 7 |
| 185 | R:R:I399 | R:R:T353 | 9.12 | No | No | 0 | 8 | 7 |
| 186 | R:R:F354 | R:R:W358 | 5.01 | Yes | Yes | 1 | 9 | 8 |
| 187 | R:R:C357 | R:R:N392 | 6.3 | No | No | 0 | 8 | 9 |
| 188 | R:R:F361 | R:R:W358 | 6.01 | Yes | Yes | 1 | 8 | 8 |
| 189 | R:R:F362 | R:R:W358 | 6.01 | Yes | Yes | 1 | 7 | 8 |
| 190 | R:R:G389 | R:R:W358 | 8.44 | No | Yes | 0 | 8 | 8 |
| 191 | R:R:N392 | R:R:W358 | 14.69 | No | Yes | 0 | 9 | 8 |
| 192 | R:R:I363 | R:R:L359 | 7.14 | No | No | 0 | 7 | 7 |
| 193 | R:R:F361 | R:R:F362 | 12.86 | Yes | Yes | 1 | 8 | 7 |
| 194 | R:R:A365 | R:R:F361 | 4.16 | No | Yes | 0 | 8 | 8 |
| 195 | R:R:F361 | R:R:I385 | 3.77 | Yes | Yes | 0 | 8 | 6 |
| 196 | R:R:F361 | R:R:N386 | 6.04 | Yes | No | 0 | 8 | 7 |
| 197 | R:R:M377 | R:R:V364 | 6.09 | Yes | No | 0 | 4 | 6 |
| 198 | R:R:L368 | R:R:M377 | 8.48 | No | Yes | 0 | 4 | 4 |
| 199 | R:R:C371 | R:R:C375 | 7.28 | No | Yes | 5 | 5 | 4 |
| 200 | R:R:M377 | R:R:P378 | 5.03 | Yes | No | 6 | 4 | 7 |
| 201 | R:R:L381 | R:R:M377 | 4.24 | No | Yes | 6 | 6 | 4 |
| 202 | R:R:L381 | R:R:P378 | 9.85 | No | No | 6 | 6 | 7 |
| 203 | R:R:I384 | R:R:L380 | 4.28 | No | No | 0 | 4 | 3 |
| 204 | R:R:I385 | R:R:L381 | 4.28 | Yes | No | 0 | 6 | 6 |
| 205 | R:R:W387 | R:R:Y390 | 16.4 | No | Yes | 0 | 8 | 8 |
| 206 | R:R:N392 | R:R:N396 | 12.26 | No | Yes | 0 | 9 | 9 |
| 207 | R:R:F403 | R:R:V398 | 3.93 | No | No | 0 | 8 | 6 |
| 208 | R:R:I399 | R:R:Y400 | 7.25 | No | Yes | 0 | 8 | 9 |
| 209 | R:R:A401 | R:R:F407 | 8.32 | No | Yes | 0 | 8 | 8 |
| 210 | R:R:F411 | R:R:Y402 | 7.22 | No | Yes | 0 | 8 | 6 |
| 211 | R:R:D406 | R:R:N404 | 5.39 | No | No | 0 | 7 | 9 |
| 212 | R:R:F407 | R:R:N404 | 7.25 | Yes | No | 0 | 8 | 9 |
| 213 | R:R:P150 | R:R:T149 | 3.5 | No | No | 0 | 5 | 8 |
| 214 | R:R:C371 | R:R:S374 | 3.44 | No | No | 5 | 5 | 5 |
| 215 | R:R:C375 | R:R:S374 | 3.44 | Yes | No | 5 | 4 | 5 |
| 216 | R:R:A50 | R:R:S86 | 3.42 | No | No | 0 | 8 | 8 |
| 217 | R:R:C119 | R:R:V85 | 3.42 | No | No | 0 | 7 | 8 |
| 218 | R:R:A79 | R:R:V57 | 3.39 | No | Yes | 0 | 9 | 9 |
| 219 | R:R:I169 | R:R:P170 | 3.39 | No | Yes | 0 | 6 | 8 |
| 220 | R:R:I385 | R:R:P360 | 3.39 | Yes | No | 0 | 6 | 9 |
| 221 | R:R:K191 | R:R:P369 | 3.35 | No | No | 0 | 5 | 7 |
| 222 | R:R:C109 | R:R:I113 | 3.27 | Yes | No | 2 | 9 | 7 |
| 223 | R:R:C187 | R:R:I113 | 3.27 | Yes | No | 2 | 9 | 7 |
| 224 | R:R:A114 | R:R:M172 | 3.22 | No | No | 0 | 7 | 6 |
| 225 | R:R:D140 | R:R:P141 | 3.22 | No | No | 0 | 7 | 8 |
| 226 | R:R:C56 | R:R:L52 | 3.17 | No | No | 0 | 8 | 4 |
| 227 | R:R:C128 | R:R:L210 | 3.17 | Yes | No | 0 | 7 | 7 |
| 228 | R:R:A94 | R:R:L43 | 3.15 | No | No | 0 | 7 | 7 |
| 229 | R:R:A50 | R:R:L394 | 3.15 | No | No | 0 | 8 | 7 |
| 230 | R:R:A55 | R:R:L83 | 3.15 | No | Yes | 0 | 5 | 8 |
| 231 | R:R:A410 | R:R:L67 | 3.15 | No | Yes | 0 | 8 | 8 |
| 232 | R:R:A153 | R:R:L74 | 3.15 | No | No | 0 | 7 | 8 |
| 233 | R:R:T196 | R:R:T200 | 3.14 | No | Yes | 0 | 7 | 7 |
| 234 | R:R:C109 | R:R:D110 | 3.11 | Yes | No | 0 | 9 | 7 |
| 235 | R:R:A410 | R:R:E64 | 3.02 | No | No | 0 | 8 | 7 |
| 236 | R:R:L67 | R:R:S66 | 3 | Yes | No | 0 | 8 | 6 |
| 237 | R:R:R152 | R:R:W136 | 3 | No | No | 0 | 7 | 5 |
| 238 | R:R:L41 | R:R:V37 | 2.98 | No | No | 0 | 4 | 5 |
| 239 | R:R:N54 | R:R:S86 | 2.98 | Yes | No | 0 | 9 | 8 |
| 240 | R:R:L83 | R:R:V87 | 2.98 | Yes | No | 0 | 8 | 6 |
| 241 | R:R:L88 | R:R:V87 | 2.98 | No | No | 0 | 6 | 6 |
| 242 | R:R:N146 | R:R:V145 | 2.96 | No | No | 0 | 5 | 6 |
| 243 | R:R:L43 | R:R:T39 | 2.95 | No | No | 0 | 7 | 5 |
| 244 | R:R:I138 | R:R:I218 | 2.94 | No | No | 0 | 9 | 9 |
| 245 | R:R:I349 | R:R:I399 | 2.94 | No | No | 0 | 7 | 8 |
| 246 | R:R:I355 | R:R:M351 | 2.92 | No | Yes | 0 | 7 | 8 |
| 247 | R:R:F370 | R:R:P369 | 2.89 | No | No | 0 | 5 | 7 |
| 248 | R:R:I38 | R:R:L42 | 2.85 | No | No | 0 | 3 | 5 |
| 249 | R:R:I414 | R:R:L63 | 2.85 | No | No | 0 | 7 | 6 |
| 250 | R:R:I142 | R:R:N146 | 2.83 | No | No | 0 | 6 | 5 |
| 251 | R:R:K147 | R:R:N146 | 2.8 | No | No | 0 | 5 | 5 |
| 252 | R:R:L394 | R:R:L46 | 2.77 | No | No | 0 | 7 | 7 |
| 253 | R:R:A93 | R:R:F112 | 2.77 | No | No | 0 | 7 | 6 |
| 254 | R:R:D110 | R:R:M172 | 2.77 | No | No | 0 | 7 | 6 |
| 255 | R:R:D406 | R:R:K405 | 2.77 | No | No | 0 | 7 | 6 |
| 256 | R:R:A137 | R:R:Y144 | 2.67 | No | No | 0 | 8 | 8 |
| 257 | R:R:F219 | R:R:T343 | 2.59 | Yes | No | 0 | 6 | 8 |
| 258 | R:R:K334 | R:R:R333 | 2.48 | No | No | 0 | 5 | 4 |
| 259 | R:R:F48 | R:R:L52 | 2.44 | No | No | 0 | 4 | 4 |
| 260 | R:R:Q68 | R:R:R65 | 2.34 | No | No | 0 | 8 | 7 |
| 261 | R:R:A79 | R:R:G76 | 1.95 | No | No | 0 | 9 | 6 |
| 262 | R:R:G105 | R:R:T108 | 1.82 | No | No | 0 | 7 | 6 |
| 263 | R:R:G382 | R:R:I385 | 1.76 | No | Yes | 0 | 4 | 6 |
| 264 | R:R:G174 | R:R:L173 | 1.71 | No | No | 0 | 5 | 6 |
| 265 | R:R:C371 | R:R:V367 | 1.71 | No | No | 5 | 5 | 6 |
| 266 | R:R:C375 | R:R:V367 | 1.71 | Yes | No | 5 | 4 | 6 |
| 267 | R:R:G389 | R:R:L388 | 1.71 | No | No | 0 | 8 | 8 |
| 268 | R:R:A79 | R:R:V58 | 1.7 | No | No | 0 | 9 | 7 |
| 269 | R:R:C49 | R:R:T45 | 1.69 | No | No | 0 | 5 | 4 |
| 270 | R:R:I47 | R:R:P91 | 1.69 | No | No | 0 | 7 | 9 |
| 271 | R:R:A221 | R:R:T139 | 1.68 | No | No | 0 | 6 | 8 |
| 272 | R:R:A336 | R:R:I226 | 1.62 | No | No | 0 | 5 | 6 |
| 273 | R:R:T39 | R:R:V98 | 1.59 | No | Yes | 0 | 5 | 6 |
| 274 | R:R:C49 | R:R:L394 | 1.59 | No | No | 0 | 5 | 7 |
| 275 | R:R:C375 | R:R:L368 | 1.59 | Yes | No | 0 | 4 | 4 |
| 276 | R:R:A59 | R:R:L63 | 1.58 | No | No | 0 | 5 | 6 |
| 277 | R:R:A336 | R:R:L337 | 1.58 | No | No | 0 | 5 | 5 |
| 278 | R:R:A71 | R:R:N69 | 1.56 | No | No | 0 | 8 | 8 |
| 279 | R:R:A329 | R:R:N328 | 1.56 | No | No | 0 | 5 | 3 |
| 280 | R:R:I38 | R:R:S34 | 1.55 | No | No | 0 | 3 | 3 |
| 281 | R:R:I350 | R:R:T346 | 1.52 | No | No | 0 | 8 | 8 |
| 282 | R:R:F219 | R:R:G216 | 1.51 | Yes | No | 0 | 6 | 3 |
| 283 | R:R:L46 | R:R:S391 | 1.5 | No | No | 0 | 7 | 5 |
| 284 | R:R:K342 | R:R:T346 | 1.5 | No | No | 0 | 8 | 8 |
| 285 | R:R:L99 | R:R:V98 | 1.49 | No | Yes | 0 | 5 | 6 |
| 286 | R:R:L380 | R:R:T379 | 1.47 | No | No | 0 | 3 | 2 |
| 287 | R:R:E372 | R:R:S373 | 1.44 | No | No | 0 | 3 | 3 |
| 288 | R:R:I169 | R:R:L173 | 1.43 | No | No | 0 | 6 | 6 |
| 289 | R:R:K332 | R:R:N328 | 1.4 | No | No | 0 | 6 | 3 |
| 290 | R:R:A155 | R:R:R151 | 1.38 | No | No | 0 | 3 | 6 |
| 291 | R:R:L159 | R:R:L162 | 1.38 | No | No | 0 | 3 | 5 |
| 292 | R:R:K405 | R:R:Q408 | 1.36 | No | No | 0 | 6 | 9 |
| 293 | R:R:A186 | R:R:Y96 | 1.33 | No | Yes | 0 | 4 | 6 |
| 294 | R:R:N100 | R:R:Q36 | 1.32 | No | No | 0 | 5 | 4 |
| 295 | R:R:F219 | R:R:V344 | 1.31 | Yes | No | 0 | 6 | 8 |
| 296 | R:R:R151 | R:R:T149 | 1.29 | No | No | 0 | 6 | 8 |
| 297 | R:R:T188 | R:R:Y96 | 1.25 | No | Yes | 0 | 5 | 6 |
| 298 | R:R:L337 | R:R:R333 | 1.21 | No | No | 0 | 5 | 4 |
| 299 | R:R:L166 | R:R:Y198 | 1.17 | No | No | 0 | 5 | 5 |
| Color | ConSurf Grade |
| No Conservation data available | |
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 | |
| 6 | |
| 7 | |
| 8 | |
| 9 |
Index: hub id, click on each number to highlight the corresponding hub in the 3D visualization.
Hub: the hub being considered.
Avg Int. Strength: the average interaction strength of all the links of the corresponding hub.
Num Of Links: the number of links of the corresponding hub.
Community: the id of the community the link belong to, otherwise 0.
ConSurf: this column reports the ConSurf conservation grades of each hub.
| Index | Hub | Avg Int. Strength | Num Of Links | Community | ConSurf |
|---|---|---|---|---|---|
| 1 | L:L:?1 | 10.9233 | 9 | 1 | 0 |
| 2 | R:R:N54 | 5.425 | 4 | 0 | 9 |
| 3 | R:R:V57 | 4.265 | 4 | 0 | 9 |
| 4 | R:R:L67 | 4.678 | 5 | 4 | 8 |
| 5 | R:R:L78 | 6.90667 | 6 | 1 | 9 |
| 6 | R:R:D82 | 7.19 | 5 | 1 | 9 |
| 7 | R:R:L83 | 3.68 | 4 | 0 | 8 |
| 8 | R:R:Y96 | 4.5475 | 4 | 2 | 6 |
| 9 | R:R:V98 | 3.8175 | 4 | 0 | 6 |
| 10 | R:R:W102 | 8.19 | 7 | 2 | 9 |
| 11 | R:R:C109 | 5.0475 | 4 | 2 | 9 |
| 12 | R:R:D116 | 10.5125 | 4 | 1 | 8 |
| 13 | R:R:T121 | 4.75 | 4 | 0 | 8 |
| 14 | R:R:W125 | 10.905 | 4 | 1 | 7 |
| 15 | R:R:H126 | 9.49571 | 7 | 1 | 8 |
| 16 | R:R:L127 | 7.09 | 5 | 1 | 8 |
| 17 | R:R:C128 | 3.93 | 4 | 1 | 7 |
| 18 | R:R:R134 | 6.42 | 4 | 1 | 9 |
| 19 | R:R:Y135 | 9.15 | 5 | 0 | 9 |
| 20 | R:R:R148 | 11.495 | 4 | 0 | 8 |
| 21 | R:R:W161 | 8.515 | 6 | 1 | 9 |
| 22 | R:R:P170 | 5.0875 | 4 | 3 | 8 |
| 23 | R:R:P171 | 6.2125 | 4 | 3 | 7 |
| 24 | R:R:C187 | 6.2575 | 4 | 2 | 9 |
| 25 | R:R:S190 | 5.625 | 4 | 3 | 6 |
| 26 | R:R:Y195 | 8.604 | 5 | 3 | 8 |
| 27 | R:R:T200 | 4.495 | 4 | 1 | 7 |
| 28 | R:R:F204 | 10.045 | 6 | 1 | 8 |
| 29 | R:R:Y205 | 6.785 | 4 | 1 | 7 |
| 30 | R:R:M211 | 5.926 | 5 | 1 | 9 |
| 31 | R:R:Y215 | 6.74571 | 7 | 1 | 9 |
| 32 | R:R:F219 | 3.094 | 5 | 0 | 6 |
| 33 | R:R:M351 | 4.37 | 4 | 1 | 8 |
| 34 | R:R:F354 | 6.20833 | 6 | 1 | 9 |
| 35 | R:R:W358 | 7.91375 | 8 | 1 | 8 |
| 36 | R:R:F361 | 9.71667 | 6 | 1 | 8 |
| 37 | R:R:F362 | 10.854 | 5 | 1 | 7 |
| 38 | R:R:C375 | 3.505 | 4 | 5 | 4 |
| 39 | R:R:M377 | 5.96 | 4 | 6 | 4 |
| 40 | R:R:I385 | 3.3 | 4 | 0 | 6 |
| 41 | R:R:Y390 | 9.71 | 6 | 1 | 8 |
| 42 | R:R:N396 | 11.6 | 4 | 1 | 9 |
| 43 | R:R:Y400 | 7.01 | 6 | 1 | 9 |
| 44 | R:R:Y402 | 5.1625 | 4 | 0 | 6 |
| 45 | R:R:F407 | 8.004 | 5 | 4 | 8 |
| Color | ConSurf Grade |
| No Conservation data available | |
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 | |
| 6 | |
| 7 | |
| 8 | |
| 9 |
Index: link id, click on each number to highlight the corresponding link in the 3D visualization.
Node1 Node2: the two nodes of the corresponding link.
Recurrence: the relative Recurrence in the pool of shortest paths.
Int. Strength: the interaction strength between the two nodes.
Hub1?, Hub2?: "Yes" if the corresponding node has more than 3 links, otherwise "No".
Community: the id of the community the link belong to, otherwise 0.
ConSurf1, ConSurf2: these columns report the ConSurf conservation grades of the two nodes involved in a link.
| Index | Node1 | Node2 | Recurrence | Int. Strength | Hub1? | Hub2? | Community | ConSurf1 | ConSurf2 |
|---|---|---|---|---|---|---|---|---|---|
| 1 | L:L:?1 | R:R:Y390 | 84.2069 | 14.37 | Yes | Yes | 1 | 0 | 8 |
| 2 | R:R:W387 | R:R:Y390 | 42.6948 | 16.4 | No | Yes | 0 | 8 | 8 |
| 3 | R:R:L43 | R:R:W387 | 34.399 | 18.22 | No | No | 0 | 7 | 8 |
| 4 | R:R:L43 | R:R:T39 | 28.8199 | 2.95 | No | No | 0 | 7 | 5 |
| 5 | R:R:T39 | R:R:V98 | 25.9674 | 1.59 | No | Yes | 0 | 5 | 6 |
| 6 | R:R:L90 | R:R:Y390 | 11.5193 | 8.21 | No | Yes | 0 | 8 | 8 |
| 7 | L:L:?1 | R:R:C120 | 40.2465 | 4.58 | Yes | No | 1 | 0 | 7 |
| 8 | R:R:C120 | R:R:W358 | 62.8247 | 5.22 | No | Yes | 1 | 7 | 8 |
| 9 | R:R:N392 | R:R:W358 | 65.1736 | 14.69 | No | Yes | 0 | 9 | 8 |
| 10 | R:R:N392 | R:R:N396 | 63.8484 | 12.26 | No | Yes | 0 | 9 | 9 |
| 11 | R:R:D82 | R:R:N396 | 63.0201 | 9.42 | Yes | Yes | 1 | 9 | 9 |
| 12 | R:R:D82 | R:R:N54 | 90.9787 | 8.08 | Yes | Yes | 0 | 9 | 9 |
| 13 | R:R:L83 | R:R:N54 | 14.7462 | 4.12 | Yes | Yes | 0 | 8 | 9 |
| 14 | L:L:?1 | R:R:F361 | 43.7152 | 25.46 | Yes | Yes | 1 | 0 | 8 |
| 15 | R:R:F361 | R:R:W358 | 48.6847 | 6.01 | Yes | Yes | 1 | 8 | 8 |
| 16 | L:L:?1 | R:R:F362 | 67.9499 | 8.78 | Yes | Yes | 1 | 0 | 7 |
| 17 | R:R:F362 | R:R:W358 | 45.7328 | 6.01 | Yes | Yes | 1 | 7 | 8 |
| 18 | R:R:N54 | R:R:P397 | 62.8512 | 6.52 | Yes | No | 0 | 9 | 9 |
| 19 | R:R:P397 | R:R:V57 | 60.923 | 3.53 | No | Yes | 0 | 9 | 9 |
| 20 | R:R:V57 | R:R:Y402 | 14.7462 | 5.05 | Yes | Yes | 0 | 9 | 6 |
| 21 | R:R:A401 | R:R:V57 | 37.2681 | 5.09 | No | Yes | 0 | 8 | 9 |
| 22 | R:R:A401 | R:R:F407 | 35.2538 | 8.32 | No | Yes | 0 | 8 | 8 |
| 23 | R:R:F407 | R:R:L67 | 16.7208 | 6.09 | Yes | Yes | 4 | 8 | 8 |
| 24 | R:R:A410 | R:R:L67 | 12.659 | 3.15 | No | Yes | 0 | 8 | 8 |
| 25 | R:R:A410 | R:R:E64 | 10.5652 | 3.02 | No | No | 0 | 8 | 7 |
| 26 | R:R:F354 | R:R:W358 | 91.6413 | 5.01 | Yes | Yes | 1 | 9 | 8 |
| 27 | R:R:F354 | R:R:M211 | 90.5546 | 11.2 | Yes | Yes | 1 | 9 | 9 |
| 28 | R:R:M211 | R:R:Y215 | 100 | 3.59 | Yes | Yes | 1 | 9 | 9 |
| 29 | R:R:I218 | R:R:Y215 | 80.3008 | 7.25 | No | Yes | 0 | 9 | 9 |
| 30 | R:R:I218 | R:R:Y135 | 76.2059 | 3.63 | No | Yes | 0 | 9 | 9 |
| 31 | R:R:W136 | R:R:Y135 | 58.8325 | 4.82 | No | Yes | 0 | 5 | 9 |
| 32 | R:R:R152 | R:R:W136 | 54.5322 | 3 | No | No | 0 | 7 | 5 |
| 33 | R:R:R148 | R:R:R152 | 36.8904 | 4.26 | Yes | No | 0 | 8 | 7 |
| 34 | R:R:R148 | R:R:V70 | 18.6523 | 14.38 | Yes | No | 0 | 8 | 7 |
| 35 | R:R:P150 | R:R:V70 | 16.3464 | 5.3 | No | No | 0 | 5 | 7 |
| 36 | R:R:F204 | R:R:F362 | 28.1441 | 21.43 | Yes | Yes | 1 | 8 | 7 |
| 37 | R:R:F204 | R:R:F354 | 33.1036 | 7.5 | Yes | Yes | 1 | 8 | 9 |
| 38 | R:R:D133 | R:R:R148 | 11.7049 | 11.91 | No | Yes | 0 | 9 | 8 |
| 39 | R:R:F354 | R:R:L127 | 34.7999 | 4.87 | Yes | Yes | 1 | 9 | 8 |
| 40 | R:R:L127 | R:R:Y400 | 14.4017 | 7.03 | Yes | Yes | 1 | 8 | 9 |
| 41 | R:R:L127 | R:R:L78 | 32.3218 | 5.54 | Yes | Yes | 1 | 8 | 9 |
| 42 | R:R:H126 | R:R:L78 | 33.6337 | 6.43 | Yes | Yes | 1 | 8 | 9 |
| 43 | R:R:L78 | R:R:N396 | 12.5696 | 12.36 | Yes | Yes | 1 | 9 | 9 |
| 44 | R:R:I130 | R:R:L78 | 15.0278 | 4.28 | No | Yes | 1 | 9 | 9 |
| 45 | R:R:H126 | R:R:W161 | 15.3359 | 5.29 | Yes | Yes | 1 | 8 | 9 |
| 46 | R:R:A93 | R:R:Y390 | 51.1198 | 4 | No | Yes | 0 | 7 | 8 |
| 47 | R:R:A93 | R:R:F112 | 48.4131 | 2.77 | No | No | 0 | 7 | 6 |
| 48 | R:R:F112 | R:R:W102 | 42.8373 | 9.02 | No | Yes | 2 | 6 | 9 |
| 49 | R:R:L99 | R:R:V98 | 14.4911 | 1.49 | No | Yes | 0 | 5 | 6 |
| 50 | R:R:C109 | R:R:W102 | 17.221 | 6.53 | Yes | Yes | 2 | 9 | 9 |
| 51 | R:R:C109 | R:R:D110 | 11.5326 | 3.11 | Yes | No | 0 | 9 | 7 |
| 52 | L:L:?1 | R:R:T121 | 18.1653 | 4.25 | Yes | Yes | 0 | 0 | 8 |
| 53 | R:R:I167 | R:R:T121 | 11.3769 | 6.08 | No | Yes | 0 | 8 | 8 |
| 54 | R:R:C128 | R:R:M211 | 13.4707 | 4.86 | Yes | Yes | 1 | 7 | 9 |
| 55 | R:R:R134 | R:R:Y400 | 15.601 | 6.17 | Yes | Yes | 1 | 9 | 9 |
| 56 | R:R:R134 | R:R:Y215 | 26.8453 | 8.23 | Yes | Yes | 1 | 9 | 9 |
| 57 | R:R:D143 | R:R:R152 | 16.3431 | 11.91 | No | No | 0 | 6 | 7 |
| 58 | L:L:?1 | R:R:I189 | 26.9215 | 7.2 | Yes | No | 0 | 0 | 6 |
| 59 | R:R:I189 | R:R:Y195 | 24.8012 | 7.25 | No | Yes | 0 | 6 | 8 |
| 60 | R:R:P170 | R:R:Y195 | 18.1918 | 5.56 | Yes | Yes | 3 | 8 | 8 |
| 61 | L:L:?1 | R:R:S199 | 27.117 | 6.49 | Yes | No | 0 | 0 | 8 |
| 62 | R:R:S199 | R:R:Y195 | 24.9006 | 11.45 | No | Yes | 0 | 8 | 8 |
| 63 | R:R:S190 | R:R:Y195 | 18.2282 | 7.63 | Yes | Yes | 3 | 6 | 8 |
| 64 | R:R:F362 | R:R:T200 | 12.1952 | 5.19 | Yes | Yes | 1 | 7 | 7 |
| 65 | R:R:T196 | R:R:T200 | 12.4006 | 3.14 | No | Yes | 0 | 7 | 7 |
| 66 | R:R:P369 | R:R:T196 | 10.8733 | 12.24 | No | No | 0 | 7 | 7 |
| 67 | R:R:L347 | R:R:Y215 | 41.2172 | 7.03 | No | Yes | 0 | 8 | 9 |
| 68 | R:R:F219 | R:R:L347 | 38.9743 | 3.65 | Yes | No | 0 | 6 | 8 |
| 69 | R:R:F219 | R:R:R223 | 27.6537 | 6.41 | Yes | No | 0 | 6 | 7 |
| 70 | R:R:R223 | R:R:R227 | 25.3711 | 13.86 | No | No | 0 | 7 | 6 |
| 71 | R:R:E340 | R:R:R227 | 23.0785 | 8.14 | No | No | 0 | 8 | 6 |
| 72 | R:R:E340 | R:R:I226 | 20.9482 | 8.2 | No | No | 0 | 8 | 6 |
| 73 | R:R:A336 | R:R:I226 | 11.7016 | 1.62 | No | No | 0 | 5 | 6 |
| 74 | R:R:I399 | R:R:Y400 | 12.3343 | 7.25 | No | Yes | 0 | 8 | 9 |
| 75 | R:R:F361 | R:R:I385 | 10.9296 | 3.77 | Yes | Yes | 0 | 8 | 6 |
| 76 | R:R:N54 | R:R:S86 | 14.7462 | 2.98 | Yes | No | 0 | 9 | 8 |
| 77 | R:R:A50 | R:R:S86 | 12.659 | 3.42 | No | No | 0 | 8 | 8 |
| 78 | R:R:I169 | R:R:P170 | 13.7126 | 3.39 | No | Yes | 0 | 6 | 8 |
| 79 | R:R:A50 | R:R:L394 | 10.5652 | 3.15 | No | No | 0 | 8 | 7 |
| 80 | R:R:D116 | R:R:Y390 | 21.2927 | 11.49 | Yes | Yes | 1 | 8 | 8 |
| 81 | R:R:C120 | R:R:D116 | 22.343 | 4.67 | No | Yes | 1 | 7 | 8 |
| 82 | R:R:D82 | R:R:L78 | 36.9997 | 8.14 | Yes | Yes | 1 | 9 | 9 |
| 83 | R:R:L127 | R:R:M211 | 26.6101 | 5.65 | Yes | Yes | 1 | 8 | 9 |
| 84 | R:R:L127 | R:R:N396 | 13.8716 | 12.36 | Yes | Yes | 1 | 8 | 9 |
| 85 | R:R:I130 | R:R:R134 | 12.7949 | 7.52 | No | Yes | 1 | 9 | 9 |
| 86 | R:R:L78 | R:R:Y400 | 14.5276 | 4.69 | Yes | Yes | 1 | 9 | 9 |
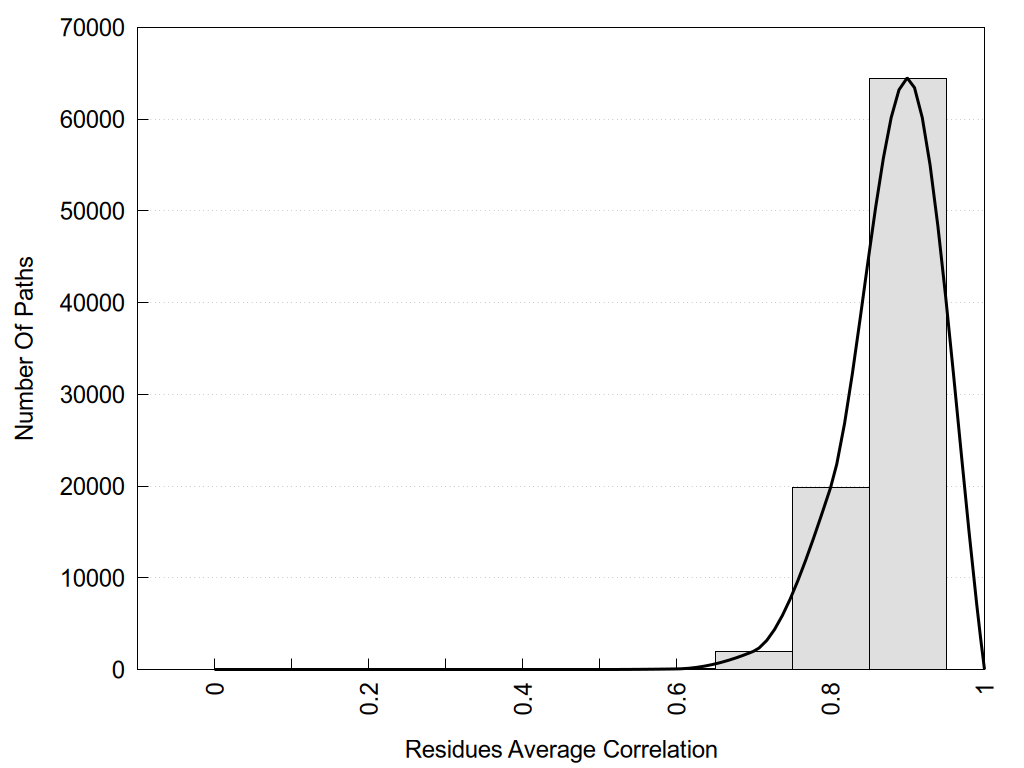
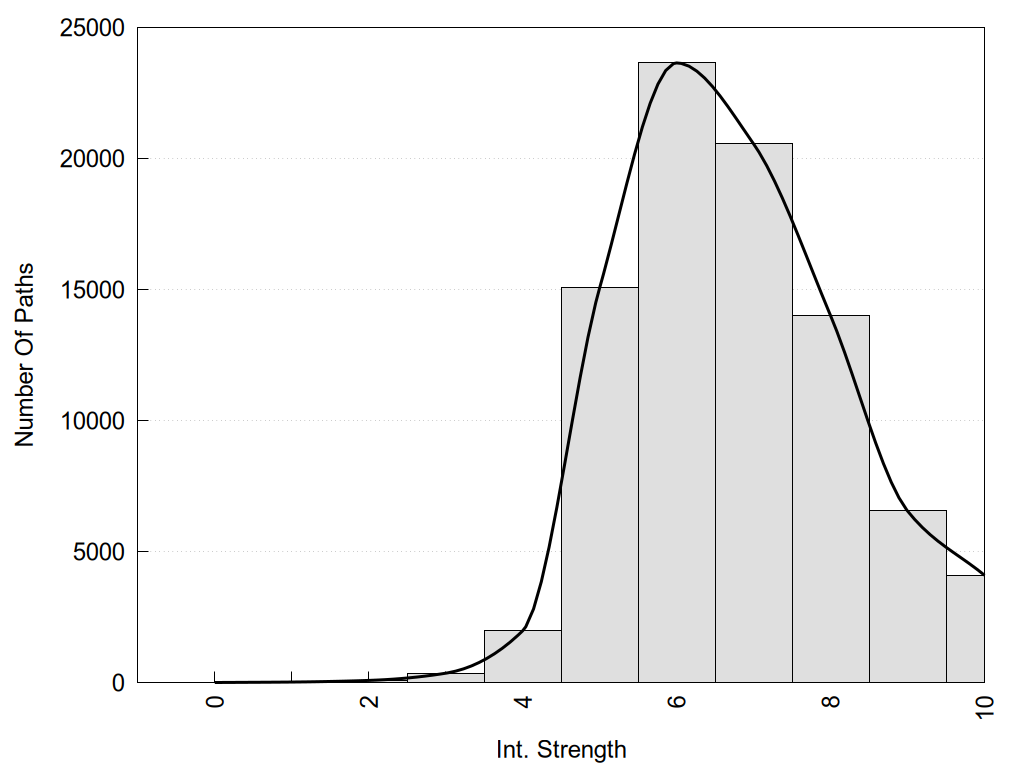
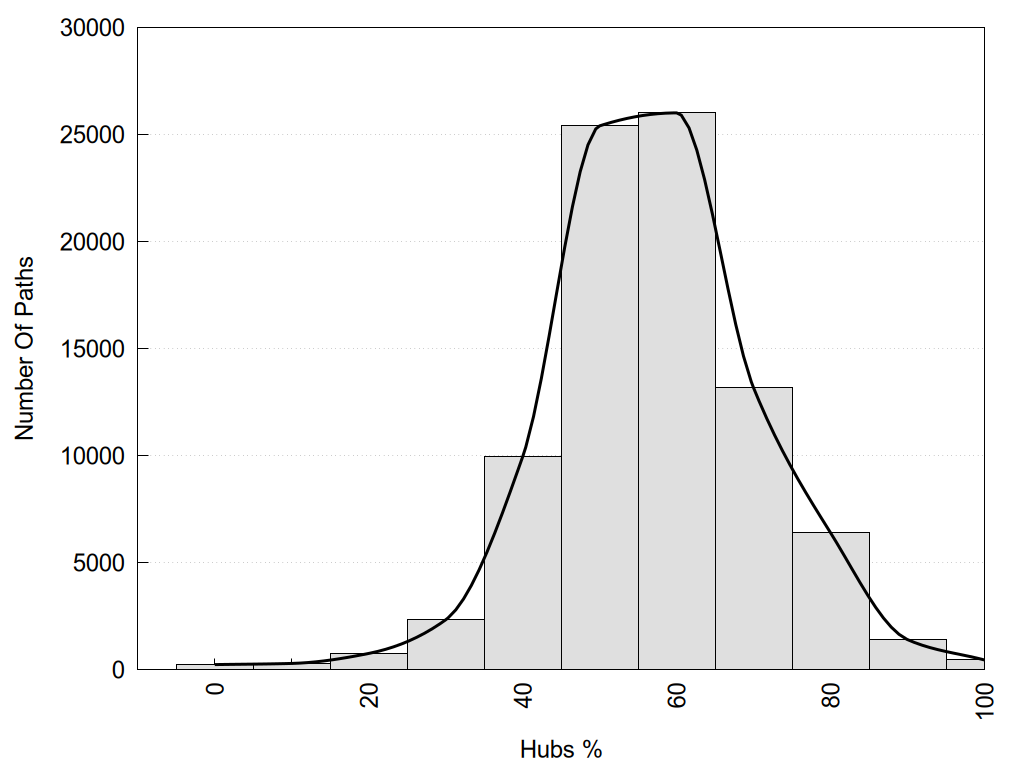
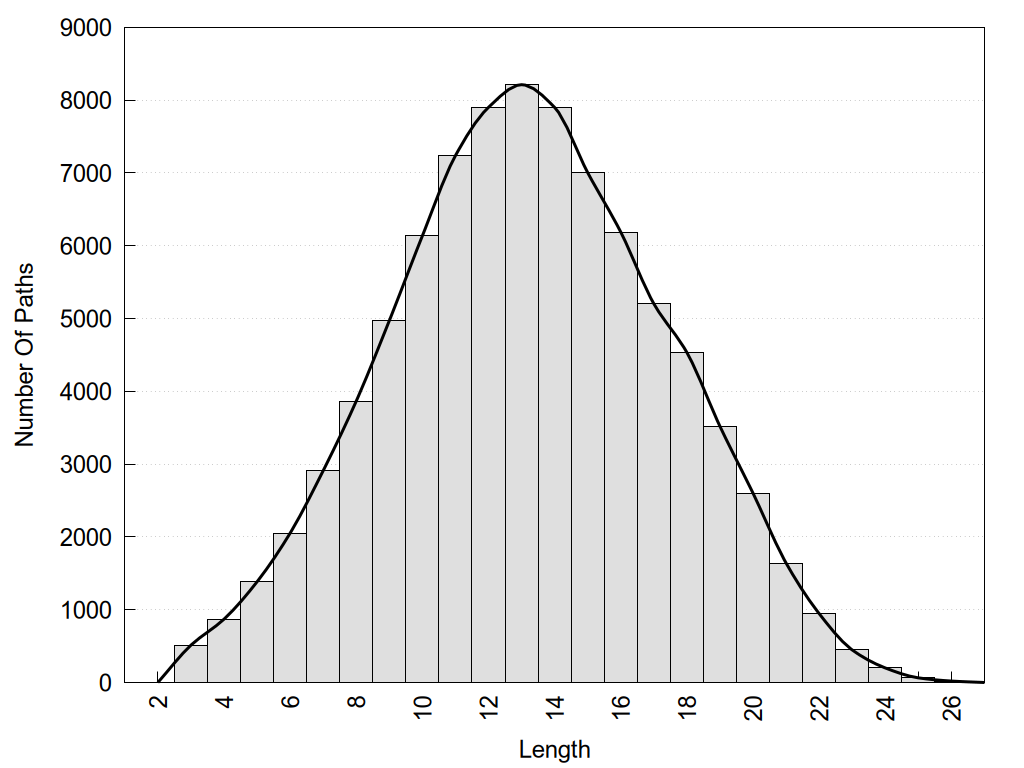
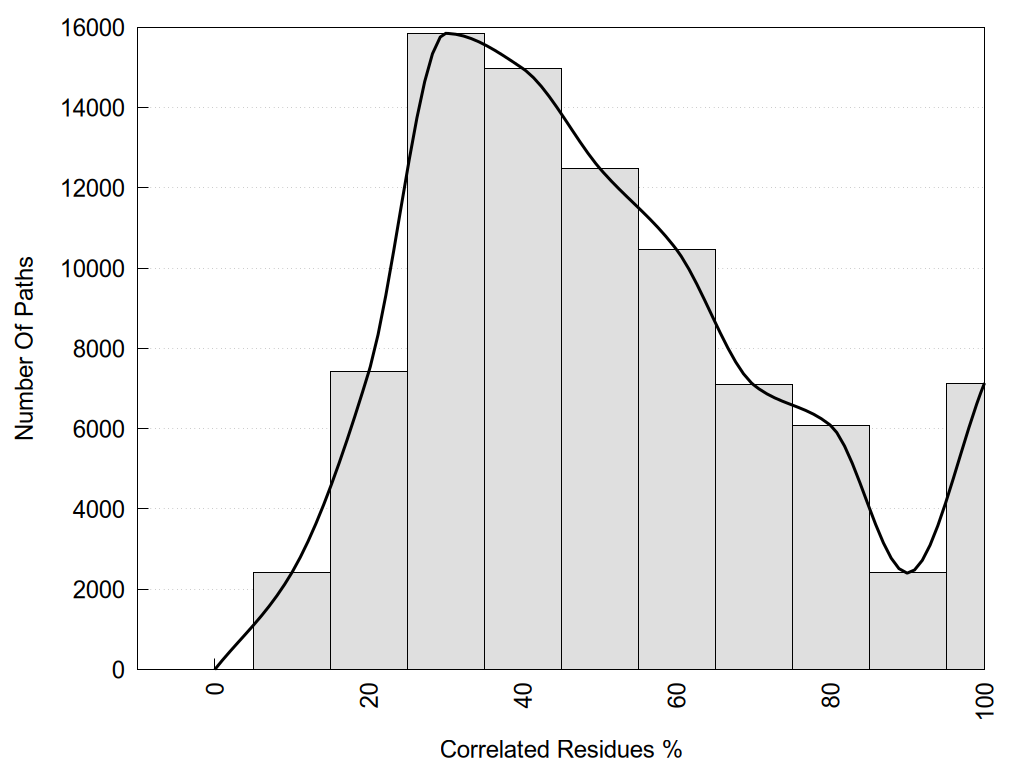
2D representation of the global metapath, ligand(s) interactions and
histograms of path distribution according to several parameters
(click on the image to enlarge it 🔍):
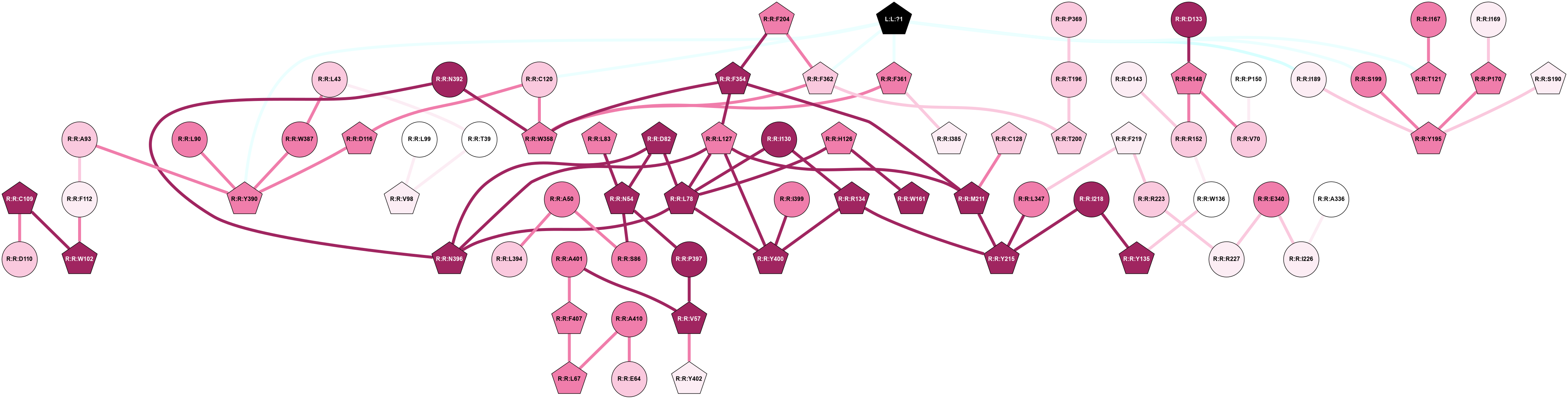
A 2D representation of the global communication in the network.
ConSurf Conservation Grade (See documentation):
n/a 1 2 3 4 5 6 7 8 9
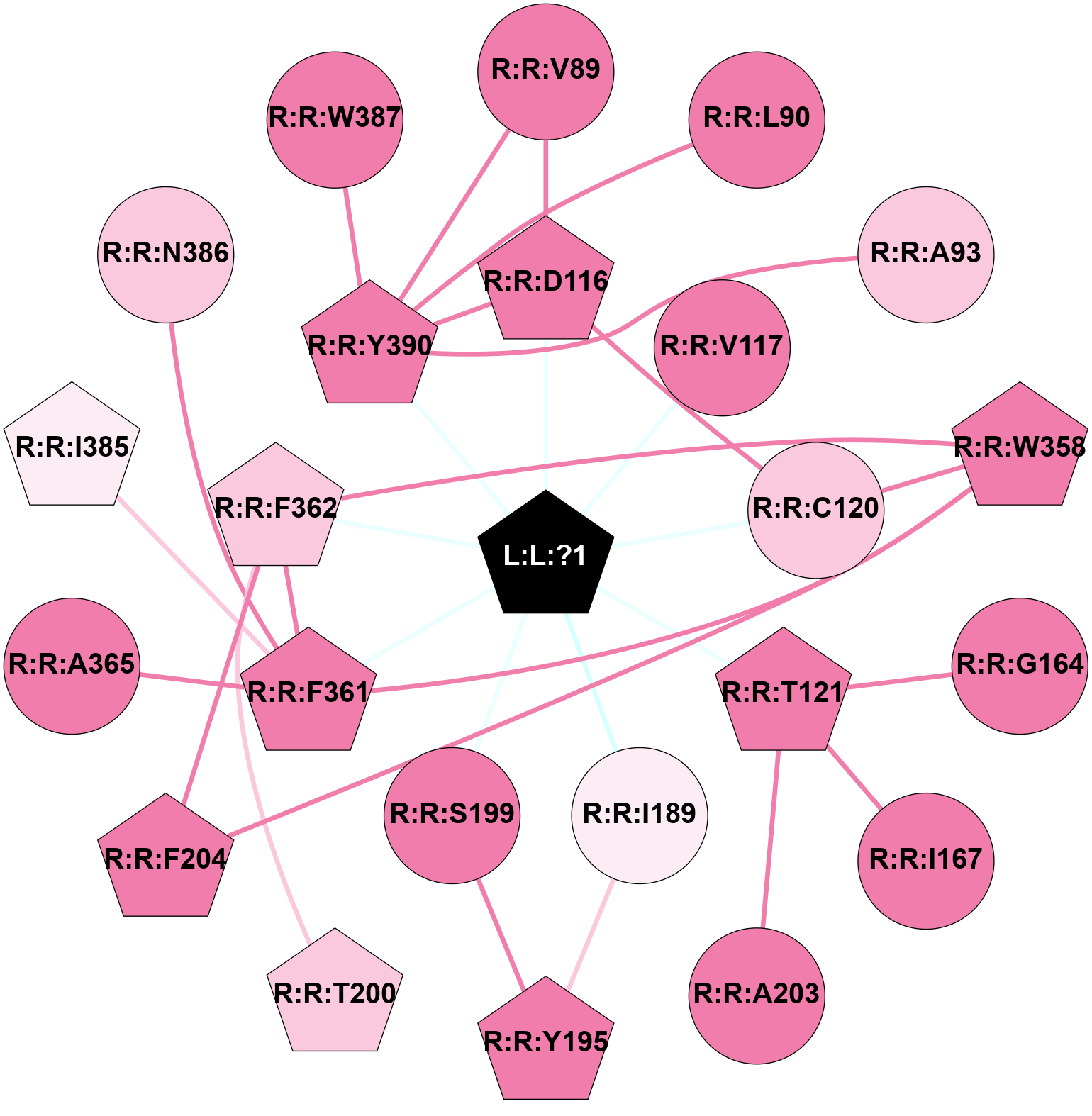
2D representation of the interactions of this orthosteric/allosteric ligand. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Links and nodes colored according to ConSurf Conservation Grade (See documentation): n/a 1 2 3 4 5 6 7 8 9 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Location and physicochemical properties of the interaction partners of this ligand | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Interactions of this ligand | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Similarities between the interactions of this ligand and those of other networks | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
Details about the values in these tables can be found in the corresponding documentation page .
| |||||||||||||||||||||||||||||||||||
| PDBsum | Open PDBsum Page |
| Chain | R |
| Protein | Receptor |
| UniProt | P08908 |
| Sequence | >9VJG_nogp_Chain_R SYQVITSLL LGTLIFCAV LGNACVVAA IALERSLQN VANYLIGSL AVTDLMVSV LVLPMAALY QVLNKWTLG QVTCDLFIA LDVLCCTSS IHLCAIALD RYWAITDPI DYVNKRTPR RAAALISLT WLIGFLISI PPMLGWRPD ACTISKDHG YTIYSTFGA FYIPLLLML VLYGRIFRA ARFRIRKTR NAEAKRKMA LARERKTVK TLGIIMGTF ILCWLPFFI VALVLPFCE SSCHMPTLL GAIINWLGY SNSLLNPVI YAYFNKDFQ NAFKKIW Click on each residue to open a popup with some information about it. ConSurf Conservation Grade (See documentation): n/a 1 2 3 4 5 6 7 8 9 |
| This receptor, from the same or other species and bound to the same or other ligands, is also present in the following networks: | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Show | PDB | Class | SubFamily | Type | SubType | Species | Orthosteric Ligand | Other Ligand(s) | Protein Partners | Resolution | Date | DOI |
| 7E2X | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | - | PtdIns4P | Gi1/β1/γ2 | 3 | 2021-04-14 | doi.org/10.1038/s41586-021-03376-8 | |
| 7E2X (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | - | PtdIns4P | 3 | 2021-04-14 | doi.org/10.1038/s41586-021-03376-8 | ||
| 7E2Y | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Serotonin | PtdIns4P | Gi1/β1/γ2 | 3 | 2021-04-14 | doi.org/10.1038/s41586-021-03376-8 | |
| 7E2Y (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Serotonin | PtdIns4P | 3 | 2021-04-14 | doi.org/10.1038/s41586-021-03376-8 | ||
| 7E2Z | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Aripiprazole | PtdIns4P | Gi1/β1/γ2 | 3.1 | 2021-04-14 | doi.org/10.1038/s41586-021-03376-8 | |
| 7E2Z (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Aripiprazole | PtdIns4P | 3.1 | 2021-04-14 | doi.org/10.1038/s41586-021-03376-8 | ||
| 8JSP | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Ulotaront | - | Gi1/β1/γ2 | 3.65 | 2023-11-15 | doi.org/10.1038/s41586-023-06804-z | |
| 8JSP (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Ulotaront | - | 3.65 | 2023-11-15 | doi.org/10.1038/s41586-023-06804-z | ||
| 8W8B | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | SEP-363856 | - | Gi1/β1/γ2 | 3 | 2023-11-22 | doi.org/10.1038/s41586-023-06775-1 | |
| 8W8B (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | SEP-363856 | - | 3 | 2023-11-22 | doi.org/10.1038/s41586-023-06775-1 | ||
| 8FY8 | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 5-MeO-DMT | PtdIns4P | Gi1/β1/γ1 | 2.79 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | |
| 8FY8 (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 5-MeO-DMT | PtdIns4P | 2.79 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | ||
| 8FYE | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 4-F,5-MeO-PyrT | PtdIns4P | Gi1/β1/γ1 | 2.85 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | |
| 8FYE (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 4-F,5-MeO-PyrT | PtdIns4P | 2.85 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | ||
| 8FYL | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Vilazodone | PtdIns4P | Gi1/β1/γ1 | 2.94 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | |
| 8FYL (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Vilazodone | PtdIns4P | 2.94 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | ||
| 8FYT | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | LSD | PtdIns4P | Gi1/β1/γ1 | 2.64 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | |
| 8FYT (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | LSD | PtdIns4P | 2.64 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | ||
| 8FYX | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | PtdIns4P | Gi1/β1/γ1 | 2.72 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | |
| 8FYX (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | PtdIns4P | 2.72 | 2024-05-15 | doi.org/10.1038/s41586-024-07403-2 | ||
| 8PJK | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | ST171 | - | Gi1/β1/γ1 | 2.4 | 2024-05-29 | doi.org/10.1126/sciadv.adv9267 | |
| 8PJK (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | ST171 | - | 2.4 | 2024-05-29 | doi.org/10.1126/sciadv.adv9267 | ||
| 8PKM | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Befiradol | - | Gi1/β1/γ1 | 2.9 | 2024-05-29 | doi.org/10.1126/sciadv.adv9267 | |
| 8PKM (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Befiradol | - | 2.9 | 2024-05-29 | doi.org/10.1126/sciadv.adv9267 | ||
| 8JT6 | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | (R)-IHCH-7179 | PtdIns4P | Gi1/β1/γ2 | 3 | 2024-02-28 | doi.org/10.1016/j.cell.2024.02.034 | |
| 8JT6 (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | (R)-IHCH-7179 | PtdIns4P | 3 | 2024-02-28 | doi.org/10.1016/j.cell.2024.02.034 | ||
| 9GL2 | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Befiradol | - | Gs/β1/γ2 | 3.2 | 2025-07-02 | doi.org/10.1126/sciadv.adv9267 | |
| 9GL2 (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Befiradol | - | 3.2 | 2025-07-02 | doi.org/10.1126/sciadv.adv9267 | ||
| 9DYD | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Asenapine | PtdIns4P | Go/β1/γ2 | 2.96 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | |
| 9DYD (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Asenapine | PtdIns4P | 2.96 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | ||
| 9DYE | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | PtdIns4P | Go/β1/γ2 | 2.9 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | |
| 9DYE (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | PtdIns4P | 2.9 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | ||
| 9DYF | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Asenapine | PtdIns4P | Gi1/β1/γ2 | 2.74 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | |
| 9DYF (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Asenapine | PtdIns4P | 2.74 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | ||
| 9MD1 | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | PtdIns4P | Gz/β1/γ2 | 3.03 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | |
| 9MD1 (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | PtdIns4P | 3.03 | 2025-08-13 | doi.org/10.1126/sciadv.adu9851 | ||
| 9VJ5 | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Gepirone | - | Go/β1/γ2 | 2.69 | 2025-11-12 | To be published | |
| 9VJ5 (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Gepirone | - | 2.69 | 2025-11-12 | To be published | ||
| 9VJ6 | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | F-15599 | - | Gi3/β1/γ2 | 2.62 | 2025-11-12 | To be published | |
| 9VJ6 (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | F-15599 | - | 2.62 | 2025-11-12 | To be published | ||
| 9VJE | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | - | Go/β1/γ2 | 2.47 | 2025-11-12 | To be published | |
| 9VJE (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | - | 2.47 | 2025-11-12 | To be published | ||
| 9VJF | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | - | Gi3/β1/γ2 | 2.7 | 2025-11-12 | To be published | |
| 9VJF (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Buspirone | - | 2.7 | 2025-11-12 | To be published | ||
| 9VJG | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 8-OH-DPAT | - | chim(NtGi2-Gz)/β1/γ2 | 2.67 | 2025-11-12 | To be published | |
| 9VJG (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 8-OH-DPAT | - | 2.67 | 2025-11-12 | To be published | ||
| 9VMY | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | TMU4142 | - | Go/β1/γ2 | 2.86 | 2025-11-12 | To be published | |
| 9VMY (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | TMU4142 | - | 2.86 | 2025-11-12 | To be published | ||
| 9VNF | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Pindolol | - | Go/β1/γ2 | 2.74 | 2025-11-12 | To be published | |
| 9VNF (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Pindolol | - | 2.74 | 2025-11-12 | To be published | ||
| 9KVG | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 5-MeO-DMT | PtdIns4P | Gi1/β1/γ2 | 2.63 | 2025-12-03 | To be published | |
| 9KVG (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | 5-MeO-DMT | PtdIns4P | 2.63 | 2025-12-03 | To be published | ||
| 9KVH | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Cinobufotenine | PtdIns4P | Gi1/β1/γ2 | 2.59 | 2025-12-03 | To be published | |
| 9KVH (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Cinobufotenine | PtdIns4P | 2.59 | 2025-12-03 | To be published | ||
| 9KVI | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Bufotenin | PtdIns4P | Gi1/β1/γ2 | 2.54 | 2025-12-03 | To be published | |
| 9KVI (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | Bufotenin | PtdIns4P | 2.54 | 2025-12-03 | To be published | ||
| 9HYI | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | DP81 | - | Gi1/β1/γ2 | 2.3 | 2026-01-21 | To be published | |
| 9HYI (No Gprot) | A | Amine | 5-Hydroxytryptamine | 5-HT1A | Homo sapiens | DP81 | - | 2.3 | 2026-01-21 | To be published | ||
You can download a compressed (zip) file with structure(s), 3D outputs (as PyMol and VMD scripts) and numerical data files (as csv and plain text files).
Download 9VJG_nogp.zipYou can click to copy the link of this page to easily come back here later
or use this QR code to link and share this page.
You can also read or download a guide explaining the meaning of all files and numerical data.